Scientists determine structure of immune molecule that counteracts HIV strains
In findings that contribute to efforts to design an AIDS vaccine, a team led by Scripps Research Institute scientists has determined the structure of an immune system antibody molecule that effectively acts against most strains of human immunodeficiency virus (HIV), the virus that causes AIDS.
The study, which is being published in an advance, online issue of the journal Proceedings of the National Academy of Sciences (PNAS) during the week of June 1, 2010, illuminates an unusual human antibody called PG16.
"This study advances the overall goal of how to design an HIV vaccine," said Scripps Research Professor Ian Wilson, who led the team with Dennis Burton, Scripps Research professor and scientific director of the International AIDS Vaccine Initiative (IAVI) Neutralizing Antibody Center at Scripps Research. "This antibody is highly effective in neutralizing HIV-1 and has evolved novel features to combat the virus."
The Problem with HIV
According to the World Health Organization's latest statistics, around 33 million people are living with HIV worldwide. During 2008 alone, more than 2 million men, women, and children succumbed to the disease and an estimated 2.7 million were infected with HIV. One of the most compelling medical challenges today is to develop a vaccine that will provide complete protection to someone who is later exposed to this virus.
HIV causes AIDS by binding to, entering, and ultimately leading to the death of T helper cells, which are immune cells that are necessary to fight off infections by common bacteria and other pathogens. As HIV depletes the body of T helper cells, common pathogens can become potentially lethal.
An effective HIV vaccine would induce antibodies (specialized immune system molecules) against the virus prior to exposure to the virus. Also called immunoglobulins, these antibodies would circulate through the blood, and track down and kill the virus.
Most of the antibodies that the body produces to fight HIV, however, are ineffective. The surface of the virus is cloaked with sugar molecules that prevent antibodies from slipping in and blocking the proteins the virus uses to latch onto a cell and infect it. To make matters more complicated, HIV is constantly mutating, so there are multiple HIV strains that antibodies elicited in any vaccine must be able to sense and destroy.
Nonetheless, while rare, broadly neutralizing antibodies against HIV do exist.
Last year, a team of scientists from IAVI, Scripps Research, Theraclone Sciences, and Monogram Biosciences published research from a systematic search for such antibodies among 2,000 volunteers. The study revealed two powerful new broadly neutralizing antibodies against HIV—PG9 and PG16, isolated from a volunteer in Africa.
"Hammerhead" Structure
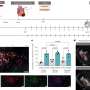
Once the broadly neutralizing antibodies were discovered, the next challenge was to figure out how they worked. To shed light on this question, in the current study members of the Wilson lab turned to x-ray crystallography, a technique that can solve structures to exquisitely high resolution.
In x-ray crystallography, scientists manipulate a protein or some other molecule so that a crystal forms. This crystal is then placed in front of a beam of x-rays, which diffract when they strike the atoms in the crystal. Based on the pattern of diffraction, scientists can reconstruct the shape of the original molecule. The scientists succeeded in forming crystals of the active part of the PG16 antibody, and in reconstructing the structure from the data—with some surprising results.
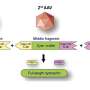
"The antibody has a novel and really interesting subdomain that hasn't been seen before," said Research Associate Rob Pejchal, who is first author of the paper. "This subdomain, which we found plays a major role in the recognition and neutralization of HIV, has a different kind of antibody architecture. We like to call it the 'hammerhead' because it resembles the head of a hammerhead shark. It reaches out from the main part of the antibody and it has two flat ends on top."
Co-author Laura Walker, a graduate student in the Scripps Research Kellogg School of Science and Technology, added, "This hypervariable loop (CDR3) that forms the novel subdomain is also unusually long for an antibody. Almost all of the antibodies we know to be broadly neutralizing against HIV have one unusual feature or another."
Pejchal notes that the study also revealed that PG16 was sulfated, suggesting possible mechanisms of action not usually seen in antibodies this effective against HIV.
While the scientists were unsuccessful so far in crystallizing PG16's sister molecule PG9, they were able to glean insight into its action from biochemical studies using both molecules. By switching a small (seven-amino acid) segment of the CDR3 subdomain of PG9 for a similar segment from PG16, the team changed the subset of HIV isolates neutralized by the antibody. This confirmed the loop in question was the "business end" of the antibody and suggested that it might be possible to create other interesting variants of the antibody by manipulating this region.
Seth Berkley, president and CEO of IAVI, which funded the study with the National Institute of Allergy and Infectious Diseases (NIAID) of the National Institute of Health (NIH), noted, "These studies of PG16 have taught us a lot about how these neutralizing antibodies work. I am particularly excited by the possibilities these findings open up for AIDS vaccine development, since the breadth and potency of HIV neutralization achieved by PG16 is what we'd like to see in the antibodies elicited by a vaccine. IAVI and its researchers will continue to support the application of these findings to the design of novel immunogens against HIV. We hope that we will be able to translate the insights gleaned from this study into the design of a promising AIDS vaccine candidate."
More information: "Crystal structure and functional studies of broadly reactive antibody PG16 reveal a novel H3 subdomain that mediates potent neutralization of HIV-1," Proceedings of the National Academy of Sciences.