New clues to how cancer-related proteins plasmin, thrombin lose inhibition
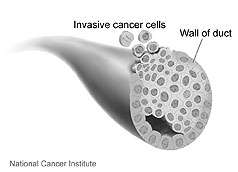
(PhysOrg.com) -- A new technique that searches blood for the tiniest remnants of broken down proteins has revealed new information about how cells crank up cancer activators called proteases. The results improve researchers' understanding of the mechanics of breast cancer and point to where to look for possible indicators of early disease.
Appearing this week in PLoS ONE, the research shows previously unknown contributing factors to protease activation, which helps spread cancer: cancer cells almost completely chew up small protein pieces that normally put the brakes on two proteases known as plasmin and thrombin. The loss of these brakes — known as protease inhibitors antiplasmin and antithrombin — occur considerably more in blood from cancer patients compared to healthy persons' blood.
Although researchers have long known that proteases become activated by cancer, this work led by researchers at the Department of Energy's Pacific Northwest National Laboratory shows two new possible mechanisms how. This work was supported by the NIH National Center for Research Resources.
Future work that measures how these proteins function in early and late stage cancer patients might reveal useful biomarkers for diagnosis.
Losing it
Cancer is largely about losing control of a cell's tightly regulated life cycle. Growing unrestrained, cancers consume bodily resources such as energy and tissue. One protein that loses control in breast and other cancers, plasmin, encourages the breakdown of the tissue matrix that keeps cells strapped together. This allows cancer cells to spread to other parts of the body. However, researchers have yet to fully work out the molecular chain of events inside and outside cells that leads to overactive plasmin.
In this work, scientists examined cellular shrapnel for clues. As cancer retunes cells to its own nefarious ends, proteins normally in use get chopped up. The cast-off proteins could reveal how cancer flourishes, providing a better understanding of the disease and insights into how to attack it.
To investigate cellular shrapnel, the team acquired blood samples from 15 breast cancer patients that spanned cancer stages from I to III. They combined all 15 samples to increase the odds of finding useful information. They also collected blood from 15 control healthy volunteers and treated the samples in the same way.
To examine just the smallest-sized cast-off protein remnants, the researchers removed the blood cells and the dozen most abundant proteins from the blood. Then they collected only the smallest protein pieces, many of which are about a tenth the size of common human proteins. The researchers called this collection of degraded pieces the plasma degradome.
Using proteomics methods at EMSL, DOE's Environmental Molecular Sciences Laboratory on the PNNL campus, the team identified the protein fragments in the cancer and healthy degradomes. The fragments between samples differed significantly. For example, cancer blood held fragments from more than 70 different proteins that were chopped into more than 800 pieces. Healthy blood only held 50 different proteins cut up into more than 400 pieces. In addition, the complement of fragments overlapped some between the cancerous and healthy, but not entirely.
Broken brakes
Different sets of fragments could indicate that cancers found new ways to cut old proteins, but an analysis showed that this was not the case. The key distinction the researchers found was in how often certain proteins were cut at individual sites — these differed tremendously. In addition, the proteins most likely to be chopped-up in the cancer samples seemed to fall into known cancer-related protein families.
"We were surprised because we expected random changes to the degradome," said lead author, PNNL biologist Yufeng Shen. "But instead we found a cluster of protein changes in very specific biological systems."
These biological systems included two proteases and their corresponding inhibitors — plasmin and antiplasmin, thrombin and antithrombin — which keep healthy cells growing properly.
When the researchers compared all the different pieces of plasmin found in the cancer samples to the healthy samples, two fragments stood out. The team found one of them, a cut-up Plg preactivation peptide, chopped seven times more often in the cancer sample than in the healthy sample. This would lead to an increase in plasmin activation in the cancer patients.
The other stand-out fragment arose from antiplasmin, and the scientists only found it in blood from cancer patients. Because antiplasmin normally prevents plasmin from destroying the environment around cells, the presence of the cut fragment meant plasmin was free to do damage.
The researchers found a similar situation with thrombin, a protease that helps blood vessels form. The cancer samples harbored fewer intact molecules of its inhibitor, antithrombin, allowing the cancer to build vessels to bring in nourishment.
AWOL Reserves
Normally, backup systems exist to deal with proteases that have gone wrong. But the scientists found evidence that the backup systems were damaged in the cancer samples as well. The team found fragments from three important backup systems: protein clusters that protect the extracellular matrix around cells; several key elements of the immune system that can scan for and kill cancer cells; and other proteins that normally suppress cells from turning cancerous.
Taken together, all these dysfunctional systems mean cells and their environment are helpless against attack by the activated proteases.
Because the team combined samples from early through late stage cancer patients, additional work is needed to determine whether either of these fragments would show up in blood before other symptoms of cancer do. If so, those fragments might serve as an early indicator of disease.
The degradome revealed many more differences than full-size proteins did. Looking through the degradome for changes between healthy and diseased samples might also provide new insights for other diseases, Shen said.
More information: Yufeng Shen, et al. "Blood Peptidome-Degradome Profile of Breast Cancer," PLoS ONE Oct. 18, 2010, DOI 10.1371/journal.pone.0013133 dx.plos.org/10.1371/journal.pone.0013133