UCLA's cancer 'roadmap' could help combat resistance to targeted drug therapies
(PhysOrg.com) -- New drugs that specifically target the mutated genes responsible for cancer growth have shown great success in extending the lives of patients, with far fewer side effects than conventional anti-cancer therapies. Unfortunately, many patients become resistant to these drugs due to secondary mutations.
Now, a multidisciplinary team of researchers at UCLA has developed a "roadmap" of the complex signaling processes involved in cancer that could lead to new methods for diagnosing and overcoming such drug resistance.
Cancer is a complicated mix of multiple, interconnected events gone awry through mutations. And while scientists have learned much about these individual events, they have long sought a better understanding of how events function together, as a system, to cause malignancies.
Proteins function as the main components of the physiological metabolic and signaling pathways of cells. Using proteomics — the large-scale study of protein interactions and activities — the UCLA team developed an approach for sorting out the complexity of events that gives rise to malignancies.
In a study published March 29 in the journal Science Signaling, the team demonstrates their use of network-scale proteomic experiments and mathematical analyses to build a "system-wide" view of how signaling mutations cause leukemia and to identify points of susceptibility that can be targeted by "cocktail" therapies to prevent drug resistance. The study is part of a journal issue focused on resistance mechanisms in cancer.
"What we need is a 'big picture' perspective," said lead author Thomas Graeber, a professor of molecular and medical pharmacology and a researcher at UCLA's Crump Institute for Molecular Imaging, Jonsson Comprehensive Cancer Center and California NanoSystems Institute. "Understanding this complex network of events is critical to designing new molecularly targeted anti-cancer therapies to simultaneously target the primary mutation while preventing the development of secondary, resistance-conferring mutations, and we now have additional tools to do this."
Molecularly targeted drugs inhibit the 'signaling' consequences of these mutational events. In many cases, drug resistance results from secondary mutations replacing or amplifying the original cancer-promoting signal targeted by the drug. The future of molecular therapies, researchers say, relies on targeting multiple events simultaneously, making it exponentially more difficult for tumor cells to develop the mutations required to escape the effect of drug therapy. This is somewhat analogous to anti-HIV drug cocktail therapies that target the inhibition of multiple viral-replication steps.
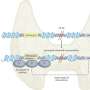
In their work, the UCLA team uses state-of-the-art technologies that concurrently measure hundreds of signaling events within cancer cells. They are trying to learn more about how these events interconnect to determine how to target the cancer cells. These new approaches for sorting out the complexities of cancer cells involve building a wiring diagram of the interconnections or "crosstalk" in cancer cells that will help scientists overcome drug resistances.
Graeber likens the goal to creating a better roadmap by identifying the bypass routes used by cancer cells to escape the inhibition caused by the drugs.
"We have the tourist information, but we need to discover the insider knowledge of the taxi driver to know how the cell gets around traffic jams rather than getting stuck in the traffic jam," he said.
"In particular, we use mass spectrometry-based proteomics to measure the activity of proteins involved in mediating the signals that cause cancer cells to grow uncontrollably," said Liudmilla Rubbi, a researcher at UCLA's Crump Institute who helped design the project. "We then analyze the resulting large-scale, quantitative data using computational algorithms to identify the informative patterns within the network of events. These patterns point us to previously unknown interactions that are critical for tumor malignancy."
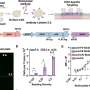
The team applied these approaches in the study of leukemias driven by the Bcr-Abl mutation, a mutation that has been successfully targeted by the well-known drug Gleevec. When drug-resistant cases develop, they become unmanageable in the clinic and are usually fatal. The researchers focused their studies on two proteins involved in clinically occurring resistance mechanisms.
An important theme that has emerged from the research is the involvement of negative feedback mechanisms in cancer growth.
"The cell has machinery to turn off cancer-promoting signaling, but typically, the effect of the mutation overwhelms the feedback mechanisms," said Björn Titz, a postdoctoral scholar and co-author of the study. "The growth of the cancer requires the correct balance between positive and negative signaling, and the state of the negative feedback mechanisms influences how the cell responds to the shutdown of the initiating mutation targeted by inhibitors such as Gleevec."
Leukemias have played a prominent role in guiding cancer research; insights discovered in leukemia research have regularly been transferred to other cancer types. The impact of Gleevec, for example, has spurred the development of other molecularly targeted anti-cancer drugs with similar modes of action.
The new roadmap also provides readout points for diagnosing patient cases that may be inherently resistant to molecularly targeted drugs, prior to any drug exposure. In the hopes of addressing this issue, Graeber's laboratory and the Crump Institute are also developing new microfluidic diagnostic technologies for making genetic and signaling measurements on tumor cells to help guide personalized medicine.
The research requires a broad range of disciplines, technology engineers and clinicians. The team includes researchers from the Crump Institute for Molecular Imaging; Institute for Molecular Medicine; Jonsson Comprehensive Cancer Center; California NanoSystems Institute; department of molecular and medical pharmacology; department of molecular, cell and developmental biology; division of rheumatology; and Institute for Genomics and Proteomics at UCLA, and from the department of laboratory medicine at the University of California, San Francisco.
The research is funded by the National Institutes of Health, the University of California Cancer Research Coordinating Committee, the Leukemia and Lymphoma Society, and the German Academic Exchange Service.