Neuro-tweets: #hashtagging the brain (w/ video)
(Medical Xpress) -- We like to think the human brain is special, something different from other brains and information processing systems, but a Cambridge professor set out to test that assumption – by conducting a live experiment using Twitter.
When he spoke at Cambridge Science Festival, Professor Ed Bullmore described how new ways of looking at the network organisation of the human brain show that it has a surprising amount in common with the worm brain, computer chips, stock markets, and many other complex systems.
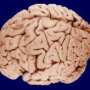
According to Professor Bullmore: “We know that the brain is fiendishly complicated in detail. Our brains have billions of nerve cells connected by trillions of synapses, so trying to figure out how it works by focusing on one cell or one synapse at a time is impossible.
“But we can use some simple mathematics to give us a different vision of the brain – losing sight of many of the details but clarifying the complex overall pattern of connections that make up a brain network.”
Viewed this way, it turns out that human brain networks represent a balance between high efficiency of information transfer and low connection cost.
Professor Bullmore discussed how different types of thinking seem to depend on different patterns of network connection and how human brains can shift rapidly between different network configurations over time. He also showed how this new research on normal brain function is beginning to change the way we think about mental health disorders, such as schizophrenia, and their treatment.
“This way of looking at the human brain tells us a lot about how it is organised in its own right. But it also allows us to ask some more general questions that might not have made sense a few years ago, such as what’s special about the human brain compared to other networks?
“You might say we are ‘taking the brain out of the skull’ to look at it directly in comparison to many superficially different information processing systems,” says Professor Bullmore.
To demonstrate this directly, Professor Bullmore conducted a live experiment during his talk at Cambridge Science Festival, the UK’s largest free science festival, as part of a collaboration between Cambridge Neuroscience and the University Communications Office.
Members of the audience and other Twitter users were asked to tweet during the lecture about the concepts that were being discussed, using the hashtag #csftwitterbrain. At the end of the talk Professor Bullmore displayed the resulting image showing the interconnectivity of the hashtagged tweets, and explained how Twitter networks can be compared to the human brain network.
“We found that the #twitterbrain network was somewhat like the brain network in being small-world and modular with highly connected hub nodes; however the brain network was more clustered and less efficient than the twitter network. So at first sight there were some points in common and some points of difference between these two information processing networks.”
“One possible explanation for the differences is that the human brain is embedded in physical space and will nearly minimize connection cost, whereas the twitter network is likely to be less constrained by the extra cost of making longer distance connections (tweets) between people.”
“It has been intriguing to see the spectacle of watching the twitter network grow or evolve over the course of several days. And I have learnt a lot about the power of new media to engage and communicate, and the potential scientific value of using Twitter to map and measure social networks.”
Brain networks video
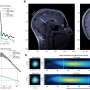
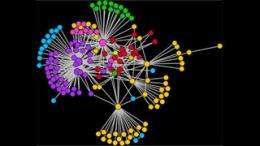
Each node of the network represents a different brain region and is colour-coded according to the larger area is located in. Pairs of nodes are linked if the activity of the two regions is found to synchronize a lot of the time during an fMRI brain scan, and the size of nodes represents how many other regions a given node is linked to.
The resulting network is used to analyze information flow in brains of healthy people as well as patients with disorders such as schizophrenia. To better understand these networks, we can decompose them into communities of nodes which are more densely connected with each other than with the rest of the network. This gives rise to a different picture, where the nodes are layed out in space according to the communities they participate in, rather than their location in real anatomical space.
The above video shows the transition from a network showing the connections between different brain regions in their anatomical locations, and a new layout emphasizing the network's structure, with nodes relocated and re-coloured based on their membership in network communities.