Gene regulator in brain's executive hub tracked across lifespan
For the first time, scientists have tracked the activity, across the lifespan, of an environmentally responsive regulatory mechanism that turns genes on and off in the brain's executive hub. Among key findings of the study by National Institutes of Health scientists: genes implicated in schizophrenia and autism turn out to be members of a select club of genes in which regulatory activity peaks during an environmentally-sensitive critical period in development. The mechanism, called DNA methylation, abruptly switches from off to on within the human brain's prefrontal cortex during this pivotal transition from fetal to postnatal life. As methylation increases, gene expression slows down after birth.
Epigenetic mechanisms like methylation leave chemical instructions that tell genes what proteins to make –what kind of tissue to produce or what functions to activate. Although not part of our DNA, these instructions are inherited from our parents. But they are also influenced by environmental factors, allowing for change throughout the lifespan.
"Developmental brain disorders may be traceable to altered methylation of genes early in life," explained Barbara Lipska, Ph.D., a scientist in the NIH's National Institute of Mental Health (NIMH) and lead author of the study. "For example, genes that code for the enzymes that carry out methylation have been implicated in schizophrenia. In the prenatal brain, these genes help to shape developing circuitry for learning, memory and other executive functions which become disturbed in the disorders. Our study reveals that methylation in a family of these genes changes dramatically during the transition from fetal to postnatal life – and that this process is influenced by methylation itself, as well as genetic variability. Regulation of these genes may be particularly sensitive to environmental influences during this critical early life period."
Lipska and colleagues report on the ebb and flow of the human prefrontal cortex's (PFC) epigenome across the lifespan, February 2, 2012, online in the American Journal of Human Genetics.
"This new study reminds us that genetic sequence is only part of the story of development. Epigenetics links nurture and nature, showing us when and where the environment can influence how the genetic sequence is read," said NIMH director Thomas R. Insel, M.D.
In a companion study published last October, the NIMH researchers traced expression of gene products in the PFC across the lifespan. The current study instead examined methylation at 27,000 sites within PFC genes that regulate such expression. Both studies examined post-mortem brains of non-psychiatrically impaired individuals ranging in age from two weeks after conception to 80 years old.
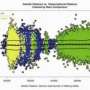
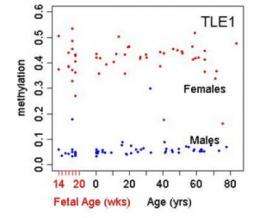
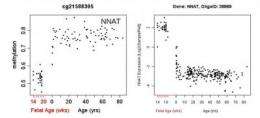
In most cases, when chemicals called methyl groups attach to regulatory regions of genes, they silence them. Usually, the more methylation, the less gene expression. Lipska's team found that the overall level of PFC methylation is low prenatally when gene expression is highest and then switches direction at birth, increasing as gene expression plummets in early childhood. It then levels off as we grow older. But methylation in some genes shows an opposite trajectory. The study found that methylation is strongly influenced by gender, age and genetic variation.
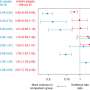
For example, methylation levels differed between males and females in 85 percent of X chromosome sites examined, which may help to explain sex differences in disorders like autism and schizophrenia.
Different genes – and subsets of genes – methylate at different ages. Some of the suspect genes found to peak in methylation around birth code for enzymes, called methytransferases, that are over-expressed in people with schizophrenia and bipolar disorder. This process is influenced, in turn, by methylation in other genes, as well as by genetic variation. So genes associated with risk for such psychiatric disorders may influence gene expression through methylation in addition to inherited DNA.
More information: Scientists worldwide can now mine a newly created online database of PFC lifespan DNA methylation from the study. The data are accessible to qualified researchers at: www.ncbi.nlm.nih.gov/projects/ … n/study.cgi?study_id = phs000417.v2.p1 . BrainCloud, a web browser application developed by NIMH to interrogate the study data, can be downloaded at BrainCloud.jhmi.edu .