Intratumor heterogeneity seen in renal carcinomas

(HealthDay) -- Extensive intratumor heterogeneity, seen in samples obtained from renal carcinomas, may lead to underestimation of the tumor genomics based on single tumor-biopsy samples, according to a study published in the March 8 issue of the New England Journal of Medicine.
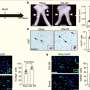
To investigate intratumor heterogeneity, Marco Gerlinger, M.D., from the Cancer Research U.K. London Research Institute, and colleagues performed exome sequencing, chromosome aberration analysis, and ploidy profiling on multiple spatially separated samples from primary renal carcinomas and associated metastatic sites. The consequences of this heterogeneity were characterized using immunohistochemical analysis, mutation functional analysis, and messenger RNA expression profiling.
The researchers found that phylogenetic reconstruction showed branched evolutionary tumor growth; and across every tumor region, 63 to 69 percent of somatic mutations were not detectable. Intratumor heterogeneity was seen for a mutation within an autoinhibitory domain of the mammalian target of rapamycin kinase, and mutational heterogeneity was observed for multiple tumor-suppressor genes converging on loss of function. In different regions of the same tumor, gene-expression signatures of good and poor prognosis were seen. Extensive intratumor heterogeneity was seen on allelic composition and ploidy profiling analysis.
"Intratumor heterogeneity may explain the difficulties encountered in the validation of oncology biomarkers owing to sampling bias, contribute to Darwinian selection of preexisting drug-resistant clones, and predict therapeutic resistance," the authors write. "Reconstructing tumor clonal architectures and the identification of common mutations located in the trunk of the phylogenetic tree may contribute to more robust biomarkers and therapeutic approaches."
The study was partially funded by Novartis; several authors disclosed financial relationships with pharmaceutical companies, including Novartis.
More information:
Full Text (subscription or payment may be required)
Editorial (subscription or payment may be required)
Copyright © 2012 HealthDay. All rights reserved.