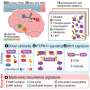
Novel type 2 diabetes genetic study involves five major ancestry groups
A consortium of scientists who are taking a novel approach in their research to detect the genetic variations that predispose individuals to type 2 diabetes provided an update of their findings at the American Society of Human Genetics (ASHG) 2012 meeting.
Among the project's novel characteristics is the ethnic diversity of the 10,000 individuals whose exomes, the 18,000 protein-coding genes, are being sequenced.
The researchers recruited 5,000 individuals with type 2 diabetes (T2D) from five major ancestry groups: African-American, East Asian, European, Hispanic and South Asian. The study population also includes an equal number of controls, individuals from these same ancestry groups who do not have T2D.
"Our hypothesis is that screening the exome in a range of diverse ethnic groups increases the range of variants of each gene surveyed, and thereby improves our ability to detect genes showing differences in the patterns of the DNA codes for proteins between individuals with type 2 diabetes and controls," said T.M. Teslovich, Ph.D., research fellow in statistical genetics at the University of Michigan, who presented the study at ASHG 2012.
The study is one of the three projects under the umbrella of the NIH-sponsored T2D-GENES (Type 2 Diabetes Genetic Exploration by Next-generation sequencing in multi-Ethnic Samples) study.
The scientists' approach also will enable them to determine whether there are T2D risk variants that are unique to an ancestry group.
An initial analysis of the data on 3,500 African-American, East Asian and South Asian individuals identified about 1.6 million single nucleotide variants (SNVs), 71.5% of which were previously unknown.
"Only about 89,000, or 5.6%, of the 1.6 million variants are present in all three groups," said Dr. Teslovich.
About 35.4% of these SNVs were unique to African-American, while 35.4% and 30.6% occurred only in East Asian and South Asian samples, respectively. Dr. Teslovich pointed out that their analysis is too preliminary to state that these population-specific variants are associated with T2D and contribute to disease risk in a single population.
By the end of 2012, the researchers will complete sequencing, which began in 2011, Dr. Teslovich said. "A total of about 5,300 individuals, half with type 2 diabetes and half controls, have been sequenced thus far," she added.
By comparing the DNA of individuals with T2D and controls, the scientists hope to isolate genes or variants that increase or reduce an individual's predisposition for developing the disease, said Dr. Teslovich.
"The unique study design will yield a catalog of variation, including alleles that are common in the population as well as those that are observed in only a small number of individuals. We'll examine each of the variants to determine which may affect an individual's risk of developing type 2 diabetes," said Dr. Teslovich.
"In addition to exome-wide analysis, we are focusing detailed mapping efforts in regions of diabetes-related traits such as fasting glucose and insulin," she added. "We anticipate that analysis of the full dataset will lead to identification of causal genes and variants."
In addition to SNVs, the researchers are searching for insertions or deletions of DNA sequence within genes as well as incorrect numbers of whole genes. The latter is referred to as copy number variations.
All the DNA sequence data and medical information will be deposited into dbGaP, the repository for genotype-phenotype relationships sponsored by the National Center for Biotechnology Information of NIH. T2D-GENES is funded by NIH's National Institute of Diabetes and Digestive and Kidney Diseases and the National Human Genome Research Institute.
A total of 75 scientists at 27 universities and other institutions are conducting T2D-GENES studies. The principal investigators of T2D-GENES are Michael Boehnke, Ph.D., University of Michigan; Mark McCarthy, M.D., University of Oxford; David Altshuler, M.D., Ph.D., Broad Institute of Harvard and MIT; Ravindranath Duggirala, Ph.D., Texas Biomedical Research Institute; and Craig Hanis Ph.D., University of Texas at Houston. Dr. McCarthy and Nancy Cox, Ph.D., University of Chicago, lead the analysis committee for this project.
More information: The researchers' abstract is titled, "Whole-exome sequencing of 10,000 type 2 diabetes cases and controls from five major ancestry groups."