Staking out unknown genomic territory

Scientists have long known that the human genome is incredibly complex. However, after almost 10 years of hard work, a team of more than 400 scientists at 32 research institutions worldwide has finally made serious headway in beginning to understand the structure, function and internal logic of the approximately 3.2 billion bases found within every cell of our body.
The Encyclopedia of DNA Elements (ENCODE) Consortium is coordinated by the US National Human Genome Research Institute and draws upon intellectual firepower from the world's leading geneticists—including Piero Carninci and colleagues at the RIKEN Omics Science Center (OSC) in Yokohama, Japan. In early September 2012, ENCODE finally shared the initial fruits of its labors with the world.
The results revealed some surprises. For example, ENCODE's most inclusive model suggests that up to 80% of the genome serves some biochemical function in at least one of the cell lines studied. ENCODE scientists also found that considerably more of the genome is dedicated to regulating gene function than to genes themselves. They have mapped many previously identified disease-associated genomic variants to such regulatory regions.
The RIKEN team was well-versed in the complexities of international collaboration through their experiences with FANTOM, a major genomics consortium headquartered at OSC, but Carninci says the guidance of lead analysis coordinator Ewan Birney was essential to the success of such an ambitious effort as ENCODE. Standardization was also a challenge, as different cells can have highly divergent patterns of gene expression. ENCODE selected 147 human cell lines and prioritized them so that all groups focused their efforts on common sets of targets.
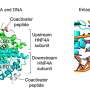
Every group had its own specialization, and Carninci and colleagues used techniques devised at RIKEN to map genome-wide sites where DNA gets transcribed into RNA2. Their team confirmed striking differences between cell lines, with no one cell type expressing more than 56.7% of the pool of RNA molecules identified in the total sample set. They also identified many cell-specific 'enhancers' of gene expression and characterized fundamental differences in expression behavior between genes that encode proteins and those that do not.
The ENCODE effort will continue but Carninci sees great value in the data already uncovered. "I believe this information will be generally used to broadly classify functional parts of the genome in many unrelated biomedical studies," he says. "We have better programs to identify regulatory elements and rules to define those elements, and can now expand this to examine, for instance, biological samples related to diseases."
More information: The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012). www.nature.com/nature/journal/ … ull/nature11247.html
Djebali, S., Davis, C.A., Merkel, A., Dobin, A., Lassmann, T., Mortazavi, A., Tanzer, A., Lagarde, J., Lin, W., Schlesinger, F. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012). www.nature.com/nature/journal/ … ull/nature11233.html