By determining structural effects of tumor-causing mutations, scientists obtain valuable information for drug discovery
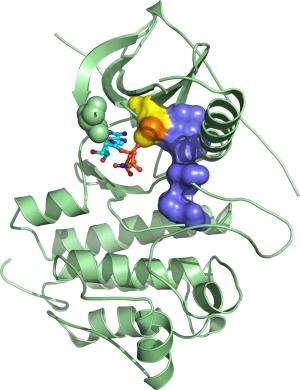
(Medical Xpress)—For many patients with non-small-cell lung carcinoma (NSCLC), tumorigenesis is fueled by mutations that hyperactivate the epidermal growth factor receptor (EGFR) signaling protein. These individuals may benefit from treatment with drugs such as gefitinib, a chemical inhibitor of EGFR, although additional mutations in EGFR can render the cancer drug-resistant.
Accordingly, scientists are struggling to overcome NSCLC recurrence. "The mutations related to drug sensitivity and those that cause drug resistance cannot be understood without the structures of these EGFR variants," explains Shigeyuki Yokoyama, director of the RIKEN Systems and Structural Biology Center in Yokohama. By teaming up with Tadashi Yamamoto's group at the University of Tokyo, Yokoyama and colleagues have now made major headway in clarifying the roots of resistance and how they might be exploited with future drugs.
The researchers performed structural and biochemical analysis of an EGFR variant containing the gefitinib sensitivity-inducing G719S mutation, either alone or with the additional resistance mutation T790M. Remarkably, they determined that although G719S binds strongly to gefitinib, G719S/T790M binds the drug even more tightly. "This appears to be contradictory to the drug resistance phenotype," says Yokoyama.
Further investigation offered potential explanations for this paradox. First, the T790M mutation appears to stabilize a network of amino acids that maintain EGFR in a continuously active state. Additionally, EGFR must bind molecules of adenosine triphosphate (ATP) to perform its signaling activities, and the researchers determined that G719S/T790M has a markedly increased capacity for ATP binding relative to G719S. This enhancement of ATP binding caused by the T790M mutation, could therefore render EGFR resistant to gefitinib in spite of its strong affinity for the drug.
Yokoyama and Yamamoto also identified the mechanistic basis for the strong drug response observed for both G719S and another common gefitinib-sensitive EGFR variant, L858R. In both cases, they identified specific rearrangements that essentially widened the protein's ATP-binding site, creating sufficient space for gefitinib to bind and interfere with signaling.
Collectively, these structural findings could prove invaluable for uncovering new vulnerabilities in drug-resistant cancers. Yokoyama and colleagues recently used computer simulations to identify vulnerabilities in the G719S/T790M double-mutant. These new data should enable even more accurate drug design against EGFR as well as other cancer-linked signaling proteins in the future. "We are planning to increase inhibitor specificity based on structure determination of the complexes between drug-resistant EGFR mutants and various compounds," says Yokoyama. "This structure-based drug discovery should yield more powerful and useful anti-cancer drugs."
More information: Yoshikawa, S., et al. Structural basis for the altered drug sensitivities of non-small cell lung cancer-associated mutants of human epidermal growth factor receptor. Oncogene 32, 27–38 (2012). www.nature.com/onc/journal/vao … full/onc201221a.html
Sato, T., et al. Identification of novel drug-resistant EGFR mutant inhibitors by in silico screening using comprehensive assessments of protein structures. Bioorganic and Medicinal Chemistry 20, 3756–3767 (2012). www.sciencedirect.com/science/ … ii/S0968089612003264