Amyloid formation: Designer proteins light the way forward
Insight into the mechanism of protein aggregation provides a model system that could lead to treatments for several associated diseases
The assembly of abnormal proteins into aggregates called amyloids is a characteristic of several diseases, including Alzheimer's, Parkinson's, diabetes type 2 and thyroid cancer. Exactly how amyloids form is unknown, but work performed at A*STAR has shown that the mechanism can be accurately recreated with specifically designed protein fragments.
The research team included Charlotte Hauser and Anupama Lakshmanan at the A*STAR Institute of Bioengineering and Nanotechnology (IBN), Daniel Cheong at the A*STAR Institute of High Performance Computing and international collaborators.
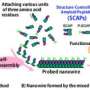
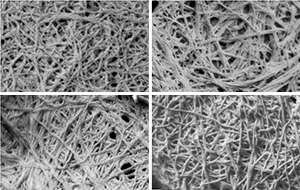
The study was based on previous work in which the IBN team designed peptides, or short protein fragments, that self-assemble in water to produce amyloids. In their recent study, the researchers used experimental techniques and computer modeling to compare the structural properties of aggregates formed by two of these peptides—LIVAGD (LD6) and IVD (ID3)—with those of peptide fragments from naturally occurring amyloid proteins (see image).
"We found that our rationally designed peptides exhibit a similar self-assembly mechanism to several amyloid-forming peptide sequences implicated in disease," says Hauser. "This provides a fresh perspective on the process of amyloid formation."
The team also aimed to clarify the role of the amino acid phenylalanine, which is often found within regions of proteins that are important for amyloid formation. Consequently, scientists thought that phenylalanine played a crucial role in the mechanism. The team's findings, however, challenge this idea: the peptides that the researchers designed contained no phenylalanine or similar amino acids, yet still formed amyloids.
Analysis of another naturally occurring peptide, KLVFFAE (KE7), reinforced this finding. KE7 is a fragment of amyloid-?, a protein involved in Alzheimer's disease. Scientists believed that two adjacent phenylalanines in its structure were crucial for amyloid formation, but Hauser and co-workers discovered that KE7 forms aggregates in a different way to the amyloid-forming peptides.
"This suggests that phenylalanine is not as essential for amyloid formation as previously postulated," explains Hauser. "It shows that there might be other core sequences that are more important."
Hauser notes that the new study forms the basis for tackling amyloid fibril formation in disease. "The fundamental mechanism of amyloid formation is believed to be common across all amyloid-related diseases, so drugs could be developed to effectively treat multiple diseases," she explains. Insufficient knowledge of protein self-assembly has hampered the search for a way to prevent or cure amyloid formation. "Our findings put forth a simplified model to study this hallmark of several degenerative disorders and design therapeutics for its control and prevention."
More information: Proceedings of the National Academy of Sciences USA 110, 519–524 (2013). www.pnas.org/content/110/2/519.abstract