New findings could help improve development of drugs for addiction
Scientists from the Florida campus of The Scripps Research Institute have described findings that could enable the development of more effective drugs for addiction with fewer side effects.
The study, published in the August 2, 2013 issue of the Journal of Biological Chemistry, showed in a combination of cell and animal studies that one active compound maintains a strong bias towards a single biological pathway, providing insight into what future drugs could look like.
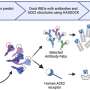
The compound examined in the study, known as 6'- guanidinonaltrindole (6'-GNTI), targets the kappa opioid receptor (KOR). Located on nerve cells, KOR plays a role in the release of dopamine, a neurotransmitter that plays a key role in drug addiction. Drugs of abuse often cause the brain to release large amounts of dopamine, flooding the brain's reward system and reinforcing the addictive cycle.
"There are a number of drug discovery efforts ongoing for KOR," said Laura Bohn, a TSRI associate professor, who led the study. "The ultimate question is how this receptor should be acted upon to achieve the best therapeutic effects. Our study identifies a marker that shows how things normally happen in live neurons—a critically important secondary test to evaluate potential compounds."
While KOR has become the focus for drug discovery efforts aimed at treating addiction and mood disorders, KOR can react to signals that originate independently from multiple biological pathways, so current drug candidates targeting KOR often produce unwanted side effects. Compounds that activate KOR can decrease the rewarding effects of abused drugs, but also induce sedation and depression.
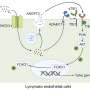
The new findings, from studies of nerve cells in the striatum (an area of the brain involved in motor activity and higher brain function), reveal a point on the KOR signaling pathway that may prove to be an important indicator of whether drug candidates can produce effects similar to the natural biological effects.
"Standard screening assays can catch differences but those differences may not play out in live tissue," Bohn noted. "Essentially, we have shown an important link between cell-based screening assays and what occurs naturally in animal models."
More information: "Functional Selectivity of 6'-guanidinonaltrindole (6'-GNTI) at Kappa Opioid Receptors in Striatal Neurons," www.jbc.org/content/early/2013 … 476234.full.pdf+html