Pathogen study explores blocking effect of E. coli O157:H7 protein

Often the key to any victory is to fully understand your opponent. This is especially true when that opponent is a significant foodborne bacterial pathogen such as E. coli O157:H7.
Philip Hardwidge, associate professor at the College of Veterinary Medicine at Kansas State University, is studying how pathogens such as E. coli use proteins to block a host's innate immune system. This system is the body's first defense against infection, often presented in the body's mucosal surfaces such as those found in the intestine.
"In terms of infectious disease, this inhibition of the human innate immune response is absolutely critical for the bacteria's ability to cause an infection," said Hardwidge, who works in the diagnostic medicine and pathobiology department. "If we can identify choke points in the interaction between the bacterium and the host, we may be able to inhibit the bacterium and prevent its survival in an infected human being."
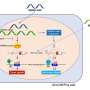
Hardwidge's lab received a multiyear grant from the National Institutes of Health to explore a protein expressed by pathogenic E. coli known as NleH1, which inhibits an important cellular signaling pathway called IKK/NF-B, or I-Kappa-Kinase/N-F-Kappa-B.
"This protein is one example of an injected bacterial protein that is able to block the innate immune system," Hardwidge said. "This protein has kind of an unusual mechanism that had not been seen in other bacterial or viral pathogens, so we're interested in understanding more about how this protein really works and whether it represents a good target for future therapeutics.
The exploration of these host-pathogen interactions requires the lab to use multidisciplinary approaches, including using animal models and advanced technologies such as quantitative polymerase chain reaction, or PCR.
"One of beauties of QPCR, or quantitative PCR, is that it gives a really reliable and easily to define comparative number of gene expression," said Mike Hays, microbiologist III in Hardwidge's lab. "It looks at a snapshot in time in that cellular environment and it could tell us at that snapshot in time, in that window, what the expression levels are of the genes that we're interested in."
Understanding how these bacterial proteins function in the host-pathogen interaction may also have applications for other human diseases.
"For example, many autoimmune diseases, many cancers and even diabetes are caused in part by an overactive component of this innate immune system," Hardwidge said. "Using information from bacteria and viruses that have evolved to block this overactive immune response, we may be able to engineer some of these bacteria proteins as potential therapeutics."
Through collaborations at Kansas State University and his position as a Chinese Academy of Sciences' senior international scientist, Hardwidge's future research will also explore both the role that the microbes that naturally live in the human body have in host-pathogen interactions and other forms of E. coli that afflict humans. Armed with this knowledge, researchers at the College of Veterinary Medicine will be able to reveal new strategies for defeating pathogens such as E. coli O157:H7.