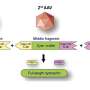
Diving deep into data to crack the gene code on disease

The key to understanding disease is in our DNA, or the human genome which contains the instructions on how our body should develop and grow.
The key to progress in genomics research is in combining as much evidence from as many different people as possible, to sort out which DNA differences (or genetic variants) are connected to particular diseases, and which are just individual variation.
We can search or mine the published biomedical literature to help scientists find this evidence, and to help interpret the significance of any given genetic change.
But in a study released this week, my colleague Antonio Jimeno Yepes and I identified a new challenge for finding evidence about genetic variants in the mass of published literature.
Much of the information is available in PubMed, the biomedical journal citation database of the National Library of Medicine in the United States and the primary repository of biomedical research.
The PubMed search tool allows researchers to search through the abstracts of published articles, and the PubMed Central resource broadens that search to full text articles, albeit for a fraction of the overall literature (2.9-million of the 22-million articles in PubMed).
So far no biomedical search tool indexes what is known as the supplementary material of journal articles, extra data associated to a paper but not part of its narrative content. It turns out this is where the vast majority of detailed evidence about individual genetic changes is located.
Lots of small genetic changes determine who we are
Take any two people, sequence their DNA, and you will typically find that they are about 99.9% identical. The remaining 0.1% determines what makes a person unique, from the shape of their ear to their risk of getting sick with a particular disease.
What seems like an insignificant amount of difference translates to millions of tiny variations at the genomic level – variations in individual bases, or the 'A', 'C', 'T', and 'G' molecules that are the building blocks of DNA.
There can be simple substitutions of one base for another, called SNPs (Singular Nucleotide Polymorphisms, pronounced "snips"), or places where one or more bases are added (insertions) or removed (deletions).
Entire segments of DNA might be repeated or moved from one place to another. Some of these changes will have no effect, and some of them will have big effects.
Sorting out which is which, and especially which changes are related to disease and how, is a major focus of modern biomedical research. Such research is possible at a level of detail and at a scale that was unimaginable not too long ago, thanks to recent improvements in DNA analysis technology called high-throughput sequencing.
Better diagnosis and treatment
Knowing which genetic changes contribute to disease risk will improve a doctor's ability to diagnose disease. It will also potentially help them to select the best treatment options, or predict the severity of the disease for a patient based on their genetic profile.
It will also lead to development of new treatments for the disease based on a deeper understanding of the underlying biology. The role of genetic variation in explaining disease is hugely important information, with the power to improve patient health.

The trouble is, those millions of tiny changes. Multiply that across even tens of people and you've got a whole heap of differences to sort through. The scale is daunting.
When you consider that it is not likely that any single change on its own will explain a person's disease risk, but rather many small changes acting together, you face an explosion of interaction combinations to explore. Biomedical researchers that are trying to make sense of all this data need some clues about where to start looking.
Finding nuggets of wisdom from the crowd
Luckily, there are plenty of researchers tackling these questions all across the globe, studying the genomes of groups of people with specific diseases and comparing them to healthy people.
When these studies identify important relationships between genetic variants and diseases, the results are usually published in a journal article. So when a researcher or clinician finds a genetic change that he or she suspects is associated with a disease, it is very possible that someone has already published some evidence that can help him confirm that suspicion.
Genome researchers spend huge amounts of time reading research publications, looking for such evidence.
But finding this evidence is hard, firstly because of just how many research publications there are to dig through. There are more than 22 million articles indexed in PubMed, and nearly a million new articles were added in 2013 alone.
Secondly, the way authors of these articles refer to genetic variants can vary tremendously. Some consistently follow the recommended nomenclature of the Human Genome Variation Society. Some authors use database identifiers, such as those from dbSNP, a large database of human SNPs. Others use more conversational ways to describe the variants, like "an adenine deletion in the mtrR promoter" [PubMed ID 23036167].
Automating the search for variants
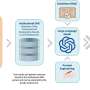
Text mining tools have been developed to automatically identify mentions of genetic variants in the published literature. These include the Extractor of Mutations (EMU) and tmVar.
Such tools have been shown to work well; they compensate for much of the different ways variants are described in text and can normalise those descriptions to the standard nomenclature.
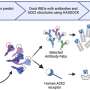
Our recent study in the journal Database, The Journal of Biological Databases and Curation applies such tools to PubMed abstracts and PubMed Central full text articles.
We aim to recover mentions of genetic variants of biological significance that have been captured in the Catalogue of Somatic Mutations in Cancer (COSMIC) database, at the Wellcome Trust Sanger Institute in the UK, and the International Society for Gastrointestinal Hereditary Tumours (InSiGHT).
We were expecting this to be a straightforward demonstration of the practical usefulness of text mining to help variant database curators and genome researchers.
We were surprised then to find that the tools only recovered less than 8% of the variants, even when we knew exactly which article to look at.
Our investigation led us to the conclusion that the vast majority of the information about genetic variants is being pulled from extra files associated with the publications, the supplementary material. When we processed these files with EMU, we were able to find more than 50% of the variants.
Broadening the search for evidence
Our study shows that the effectiveness of automatic literature mining methods for finding information about genetic variants hinges on access to the supplementary material.
This material is unfortunately not systematically indexed. Every publisher has a different way of storing and linking to the additional data files from publications. This diversity makes it difficult for automated tools to locate the data for processing.
Even if we could find all the relevant data files, there remain challenges.
When we considered the supplementary material, the EMU tool still missed about 50% of the curated variants. These are mostly variants that are included in tables or supplementary material in a way that is different from how they are typically referred to in text, such as when the information is spread across columns in a spreadsheet.
Our usual text mining tools don't work well for data expressed in this way, so new strategies are needed to automatically extract and normalise the data presented in tabular form.
Finding the genetic variants in the published literature is only the starting point for the important work of understanding the biological role of those variants, and how they are connected to a disease. The results in those publications need to be interpreted and synthesised.
But given how many publications are coming out every day, the lack of consistency in how genetic variants are described, and the fact that the information often isn't even in the papers themselves, it is clear that genome researchers need effective text mining technology to help them reach that starting point.
We are still working to get them there.
This story is published courtesy of The Conversation (under Creative Commons-Attribution/No derivatives).
![]()