How diabetes drugs may work against cancer
(Medical Xpress)—For several years, a class of anti-diabetic drugs known as biguanides, has been associated with anti-cancer properties. A number of retrospective studies have shown that the widely used diabetes drug metformin can benefit some cancer patients. Despite this intriguing correlation, it has been unclear how metformin might exert its anti-cancer effects and, perhaps more importantly, in which patients.
Now, Whitehead Institute scientists are beginning to unravel this mystery, identifying a major mitochondrial pathway that imbues cancer cells with the ability to survive in low-glucose environments. By finding cancer cells with defects in this pathway or with impaired glucose utilization, the scientists can predict which tumor will be sensitive to the anti-diabetic drugs known to inhibit the pathway in question. Their work is described online this week in the journal Nature.
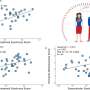
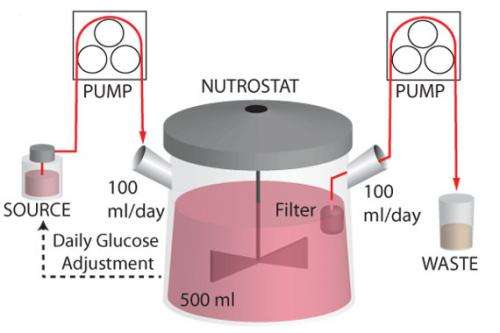
To study how cancer cells survive in the kind of low-glucose environment found within cancerous tumors, Kıvanç Birsoy and Richard Possemato, postdoctoral researchers in Whitehead Member David Sabatini's lab, developed a system that circulates low-nutrient media continuously around cells. Of the 30 cancer cell lines tested within this system, most appeared unaffected by a lack of glucose. However, a few of the lines thrived and reproduced rapidly, while others struggled. The varied responses to a glucose shortage were puzzling.
"No one really understood why cancer cells had these responses or whether they were important for the formation of the tumor," says Possemato, who coauthored the Nature paper with Birsoy. "The cancer-relevance of the alterations that we found as underlying this response to low glucose will still need to be investigated."
Birsoy and Possemato wondered whether certain cancer cells' susceptibility to a low glucose environment could be exploited to attack tumors. They screened overly distressed cells for genes whose suppression improved or further hindered the cells' survival rates. The screen flagged genes involved in glucose transportation and oxidative phosphorylation, a metabolic pathway in mitochondria. The powerhouses of a cell, mitochondria are membrane-bound organelles with their own DNA, including genes that control oxidative phosphorylation.
Birsoy and Possemato hypothesized that cancer cells with mutations in these genes are over-taxing their mitochondria under normal conditions. When placed in a harsh, low-glucose environment, the mitochondria are maxed out, and the cells suffer. If true, the hypothesis would suggest that further impairing mitochondrial function, with biguanides—which are known oxidative phosphorylation inhibitors—could push the mitochondria beyond their limits, to the detriment of the cancer cells.
They first tested their hypothesis in vitro on 13 cell lines with glucose utilization defects and mitochondrial DNA mutations. Compared to control cells, those sensitive to low glucose were five to 20 times more susceptible to phenformin, a more potent biguanide than metformin. Birsoy and Possemato then tested phenformin's effectiveness in mice implanted with tumors derived from low-glucose-sensitive cancer cells. The drug inhibited the tumors' growth.
"These results show that mitochondrial DNA mutations and glucose import defects can be used as biomarkers for biguanide sensitivity to determine if a cancer patient might benefit from these drugs," says Birsoy. "And this is the first time that anyone has shown that the direct cytotoxic effects of this class of drugs, including metformin and phenformin, on cancer cells are mediated through their effect on mitochondria."
To confirm the accuracy of their proposed biomarkers, Birsoy and Possemato want to analyze previous clinical trials to see if cancer patients with the proposed biomarkers fared better with metformin treatment than patients without the biomarkers.
More information: Kıvanç Birsoy, et al."Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides." Nature, March 16, 2014. DOI: 10.1038/nature13110