Splice variants reveal connections among autism genes
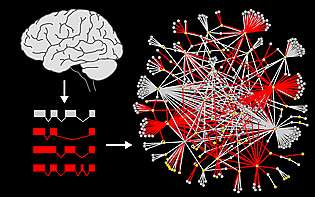
A team of researchers from the University of California, San Diego School of Medicine and the Center for Cancer Systems Biology (CCSB) at the Dana-Farber Cancer Institute has uncovered a new aspect of autism, revealing that proteins involved in autism interact with many more partners than previously known. These interactions had not been detected earlier because they involve alternatively spliced forms of autism genes found in the brain.
In their study, published in the April 11, 2014 online issue of Nature Communications, the scientists isolated hundreds of new variants of autism genes from the human brain, and then screened their protein products against thousands of other proteins to identify interacting partners. Proteins produced by alternatively-spliced autism genes and their many partners formed a biological network that produced an unprecedented view of how autism genes are connected.
"When the newly discovered splice forms of autism genes were added to the network, the total number of interactions doubled," said principal investigator Lilia Iakoucheva, PhD, assistant professor in the Department of Psychiatry at UC San Diego. In some cases, the splice forms interacted with a completely different set of proteins. "What we see from this network is that different variants of the same protein could alter the wiring of the entire system," Iakoucheva said.
"This is the first proteome-scale interaction network to incorporate alternative splice forms," noted Marc Vidal, PhD, CCSB director and a co-investigator on the study. "The fact that protein variants produce such diverse patterns of interactions is exciting and quite unexpected."
The new network also illuminated how multiple autism genes connect to one another. The scientists found that one class of mutations involved in autism, known as copy number variants, involve genes that are closely connected to each other directly or indirectly through a common partner. "This suggests that shared biological pathways may be disrupted in patients with different autism mutations," said co-first author Guan Ning Lin, PhD, a postdoctoral fellow in Iakoucheva's laboratory.
Beyond providing greater breadth and depth around autism proteins, the network represents a new resource for future autism studies, according to Iakoucheva. For example, she said the physical collection of more than 400 splicing variants of autism candidate genes could be used by other researchers interested in studying a specific protein variant. Some of the highly connected network partners may also represent potential drug targets. All interaction data will reside in the publicly available National Database of Autism Research.
"With this assembled autism network, we can begin to investigate how newly discovered mutations from patients may disrupt this network," said Iakoucheva. "This is an important task because the mechanism by which mutant proteins contribute to autism in 99.9 percent of cases remains unknown."