How breast cancer 'expresses itself'

About one in eight women in the United States will contract breast cancer in her lifetime. Now new research from Tel Aviv University-affiliated researchers, in collaboration with Johns Hopkins University, has provided another tool to help women, clinicians, and scientists searching for a cure to the one of the most widespread yet incurable diseases on the planet.
Dr. Ella Evron and Dr. Ayelet Avraham of the TAU-affiliated Assaf Harofeh Medical Center, together with Prof. Saraswati Sukumar of Johns Hopkins, have found that "gene regulation," the process that shuts off certain parts of a cell's DNA code or blueprint in healthy breast tissue cells, may also play a critical role in the development of breast cancer. Their research, published in PLOS ONE, focused on one particular gene—TRIM29—selected from a pool of 100 genes with regulatory patterns specific to normal breast tissue, to prove the link between breast-specific genes and the pathology of cancer.
"We found that normal tissue affects the cancer that grows in that organ—in other words, the specific pattern of gene regulation in the normal breast affects breast cancer, the characteristics of the disease, and its clinical behavior," said Dr. Avraham, a biologist and a researcher in the lab. "We hope that this study will lead to a better understanding of the cancer predisposition of mammary tissues and point to new targets for cancer intervention."
Searching for the right gene
In the study, normal tissue samples taken from conventional breast reduction surgeries were examined in a laboratory. The researchers isolated the milk ducts and purified the breast-tissue cells to create a cell culture, which was then tested for different gene regulation profiles.
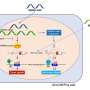
While all cell types share the same genetic code (DNA), certain genes are specifically "expressed" or "silenced" in each cell type. Consequently, the unique gene expression patterns in every tissue dictate its structure and function. Various "gatekeeper" mechanisms either allow or block gene expression in our cells. One such mechanism is "DNA methylation," which shuts off or silences parts of the genetic code to form a specific pattern that identifies each tissue type.
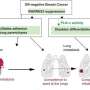
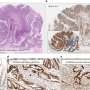
The researchers compared the DNA methylation profiles of thousands of genes in breast, colon, lung, and endometrial tissues, selecting one gene, TRIM29, for further analysis. They found that the TRIM29 gene bore a unique DNA regulation in normal and cancerous breast tissues as opposed to other organ tissues.
"In breast tissue we found that this gene was expressed in normal cells and silenced in the cancer cells," said Dr. Avraham. "In contrast, in other bodily tissues, the gene was silenced in normal cells and over-expressed in tumors. This emphasizes the link between tissue-specific gene regulation and the development of cancer."
Silencing the cancer
"Tissue-specific genes often take part in carcinogenesis. A well known example is estrogen, which is involved in the normal differentiation of the breast and also in breast cancer development," said Dr. Evron, a senior oncologist and a researcher in the lab. "Thus, the estrogen receptor over-expresses in nearly 70% of breast cancers. It's the target of very effective anti breast cancer therapy. In this study we identify more genes that have specific regulation in normal breast tissue as compared with other organ tissues."
"Another example is women who carry a mutation in the BRCA1/2 genes and develop cancers almost exclusively in the breast and ovary," said Dr. Evron. "This led to the hypothesis that these tissues are 'marked' for cancer predisposition during differentiation. Searching for these marks may throw light on this process.
"Certainly the concept of looking for genes that are involved in both in differentiation and in carcinogenesis is promising, and the novel list of breast-specific regulated genes we found may encourage further study in this direction. But we can't stop here. If we know which genes are responsible for breast cancer, then we can tailor therapies to target those genes specifically."