Innovative 'genotype first' approach uncovers protective factor for heart disease
Extensive sequencing of DNA from thousands of individuals in Finland has unearthed scores of mutations that destroy gene function and are found at unusually high frequencies. Among these are two mutations in a gene called LPA that may reduce a person's risk of heart disease. These findings are an exciting proof-of-concept for a new "genotype first" approach to identifying rare genetic variants associated with, or protecting from, disease followed by extensive medical review of carriers. The new study by researchers from the Broad Institute, Massachusetts General Hospital (MGH), the University of Helsinki, and an international team of collaborators appears in a paper published online July 31 in PLOS Genetics.
The researchers studied exomes—the portions of the genome that correspond to protein-coding genes—from 3,000 Finns and compared them to those of 3,000 non-Finnish Europeans. They identified 83 gene-deactivating variants that were at least twice as prevalent in Finns and went on to study these variants in over 35,000 Finns. Recent examples in heart disease, HIV, type 2 diabetes and Crohn's disease have demonstrated that such mutations – known as "loss-of-function" mutations – in some cases protect from, rather than cause, disease and thereby suggest new paths toward therapeutics.
Geneticists have known that Mendelian, recessive genetic diseases –such as Tay-Sachs or cystic fibrosis that are caused by a single, mutated gene – are more common in isolated populations because of a phenomenon known as "bottlenecking." When a small population is isolated for tens to hundreds of generations, the population's genetic diversity becomes restricted, and occasional rare genetic variations can by chance become much more common. While this has long been recognized as the source of the unique rare disease patterns seen in isolated populations, this paper demonstrates that the same principles can help researchers identify rare, loss-of-function variants in genome-wide association studies on these isolated populations.
In the current study, researchers chose to study modern Finns – a population that descended from a well-documented bottleneck that occurred around 4,000 years ago. Comparing Finns with their non-Finnish European counterparts gave the researchers strong, empirical data.
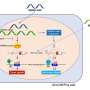
The LPA gene encodes Lipoprotein(a), a type of lipoprotein, first identified in 1963 and a known risk factor for heart disease. The variants described in this paper reduced levels of LPA gene expression causing lower levels of Lipoprotein(a) in the blood. The research team examined Finnish medical records and found that the loss-of-function variants were not associated with other health problems, making blocking LPA expression a potentially exciting therapeutic approach. The availability of centralized medical records available in Finland enabled the researchers to shift the paradigm of medical genetics to a "genotype first" approach.
"This new approach could significantly change how researchers analyze rare variants for complex diseases. It gives us a window into the genetics of complex diseases that we haven't had before," said co-senior author Mark Daly, co-director of the Program in Medical and Population Genetics at the Broad Institute and chief of the Analytic and Translational Genetics Unit for the Center for Human Genetic Research at MGH. "By combining the information from detailed medical records with the information contained in the genomes of a bottlenecked population, we're uncovering rare variants that contribute to complex diseases."
Heart disease is a leading killer globally. The World Health Organization reports that cardiovascular disease was responsible for 30 percent of all global deaths, or 17.3 million people in 2008. Therapeutics able to specifically address this risk by targeting LPA could have a global impact on medical outcomes.
This work highlights the potential for using rare variant analysis in isolated populations to study complex diseases, an approach that had previously been largely limited to Mendelian traits. The approach can now be applied to other complex diseases that have many contributing genetic factors.
"We've illustrated the validity of this approach by identifying rare, loss-of-function variants with promising therapeutic potential for the treatment of heart disease, but this work also represents a reproducible approach that can be used to increase our understanding of other complex diseases as well," said co-senior author Aarno Palotie (Broad Institute, Massachusetts General Hospital, Harvard Medical School, Institute for Molecular Medicine Finland FIMM, University of Helsinki).
More information: PLoS Genetics, Online July 31, 2014. DOI: 10.1371/journal.pgen.1004494