Targeted next-generation sequencing reveals a high number of genomic mutations in advanced malignant
Next generation sequencing in malignant pleural mesothelioma (MPM) tumors shows a complex mutational setting with a high number of genetic alterations in genes involved in DNA repair, cell survival and cell proliferation pathways. Increased accumulation of mutations correlates with early progression of the tumor and decreased survival.
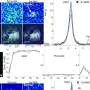
MPM is a fatal cancer associated with asbestos exposure that develops on the outer linings of the lungs. The 3-year survival rate is only 8% and most MPM patients are diagnosed with late stage disease with limited therapeutic options. Next-generation sequencing (NGS) is a robust method that determines the identity and order of nucleotides within genomic DNA molecules isolated from cells and targeted NGS focuses on a specific set of genes instead of sequencing the entire genome. The use of NGS has the potential to accurately describe the type, location, and number of genetic mutations in MPM and allow for associations with patient characteristics, including survival data.
Researchers from University of Torino, Orbassano, Italy and Saint Antonio and Biagio General Hospital, Alessandria, Italy used targeted NGS to retrospectively sequence 52 genes in 123 patients with advanced (stage III and IV) MPM and complete clinical information. Sequencing was conducted on genomic DNA isolated from archival formalin-fixed paraffin-embedded (FFPE) tumor tissue. Sanger sequencing was used to validate the NGS results and immunohistochemistry was used to examine the protein expression for two of the genes.
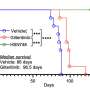
Results published in the Journal of Thoracic Oncology, the official journal of the International Association for the Study of Lung Cancer (IASLC), show that the mutations clustered in two main pathways, p53/DNA repair and PI3K-AKT. Certain mutations within the PIK3CA, STK11 or TP53 genes associated with a decreased time to disease progression. Additionally, there was a decrease in the time to disease progression and overall survival when there was an accumulation of multiple mutations. Finally, genomic mutation in the BAP1 gene, which resulted in amino acids changes within the BAP1 protein, correlated with nuclear localization of the BAP1 protein.
"Our results show that NGS is clearly feasible despite the very well-known challenges of fragmented and low-yield genomic DNA isolated from FFPE tumor tissue", said the authors. "Our data did not identify any specific mutation as a single driver gene, as found in adenocarcinoma of the lung, however our data suggests that in MPM there is an accumulation of several non-driver mutations, which may explain the extremely long latency phase of this asbestos-related disease." For the future the authors suggest that "some of the mutations will further be assessed for functional changes and could be carefully considered as stratification factors for future clinical trials investigating the role of targeted-therapies in MPM".
More information: Journal of Thoracic Oncology DOI: 10.1097/JTO.0000000000000436