April 3, 2015 report
Toward a model of synchrony in brain networks
(MedicalXpress)—Resting state networks (RSNs) in the brain are topographies of neural structures between which lag states propagate due to fluctuations of physical and other activities. Studying these networks reveals information about the functional connectivity of neural structures and regions. Results from various studies have confirmed that brain activity is spatially structured, linked to the representation of function, and has clinical relevance.
Functional connectivity is different from the brain's structural connectivity, which describes brain regions that are anatomically attached to each other. Regions with no structural connectivity can nonetheless have functional connectivity as nodes in a functionally connected RSN. Many common RSNs have been mapped in healthy subjects, and researchers believe that understanding the relationships between these networks can contribute to a fundamental model of brain function.
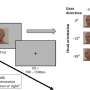
One of the tremendous advantages of functional magnetic resonance imaging (fMRI) is the ability to study brain functional activity without the need for subjects to perform complex tasks. Using fMRI to study resting-state functional connectivity yields a wealth of information about different stages of consciousness and patterns of synchronous activity. One of the neurological features that has emerged from such research is the existence of lags in intrinsic activity as represented by fluctuations of the blood-oxygen level-dependent signals (BOLDs), which are temporally synchronous within the somatomotor system.
Last year, researchers at the departments of radiology and neurology at Washington University published an analysis demonstrating that, contrary to the belief that BOLDs were synchronous with resting state networks (RSNs), the lag topography of BOLDs and RSNs is actually orthogonal. Additionally, they established that BOLDs are not attributable to hemodynamic factors and have neural origin.
In a new study published in the Proceedings of the National Academy of Sciences, the same researchers demonstrate that the propagated activity of lag threads in the brain is unidirectional within conventionally understood RSNs. Additionally, identifiable resting-state networks have been demonstrated to to emerge naturally as an occurrence of shared patterns of propagation.
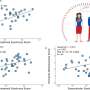
First, the study demonstrated the existence of eight separate, reproducible orthogonal lag processes across data gathered from 1,376 fMRI subjects. Drawing terminology from modern computer programming practices, the researchers refer to these lag processes as "threads," by analogy to applications with multiple independent thread sequences.
The researchers recover the lag processes in multidimensional time series using a technique called principal component analysis. They determined the sources and destinations of propagated BOLD activity and a range of lag values of ~2 seconds. Although specific anatomical structures were often the sources or the destinations of propagated threads, those paths did not respect the boundaries of RSNs: Rather, BOLD fMRI signals propagate both within and across identified RSNs.
Finding a similarity in the lag-thread correlations and the BOLD fMRI zero-lag correlations, the researchers hypothesize the existence of lag-thread motifs: timed sequences of propagation through brain regions that are shared by multiple threads. They propose that temporal synchrony between RSNs naturally emerges as a consequence of these preprogrammed motifs, and conclude that lag threads represent a fundamental organizing property of the brain's intrinsic activity.
The brain's spontaneous activity presents two seemingly contradictory behaviors. A perfectly synchronous system would have no lags, and a system with a set of lags is not synchronous. This might be a property of the brain's dual roles of segregation and integration. Neurologists are strongly interested in establishing how segregated regions of the brain become functionally integrated. The WU researchers believe that lag threads can explain how spatially segregated networks can be integrated over a time scale of seconds.
The researchers speculate about a problem posed by lag threads: How does synchrony arise if spontaneous activity is characterized by a lag structure? "Our results suggest that lag thread motifs provide an answer," they write. "Preservation of lag sequencing within certain regions of the brain (i.e., RSNs) across multiple threads gives rise to zero-lag synchrony (spatial segregation) and lags (temporal integration) in the brain's activity."
More information: "Lag threads organize the brain's intrinsic activity." PNAS 2015 ; published ahead of print March 30, 2015, DOI: 10.1073/pnas.1503960112
Abstract
It has been widely reported that intrinsic brain activity, in a variety of animals including humans, is spatiotemporally structured. Specifically, propagated slow activity has been repeatedly demonstrated in animals. In human resting-state fMRI, spontaneous activity has been understood predominantly in terms of zero-lag temporal synchrony within widely distributed functional systems (resting-state networks). Here, we use resting-state fMRI from 1,376 normal, young adults to demonstrate that multiple, highly reproducible, temporal sequences of propagated activity, which we term "lag threads," are present in the brain. Moreover, this propagated activity is largely unidirectional within conventionally understood resting-state networks. Modeling experiments show that resting-state networks naturally emerge as a consequence of shared patterns of propagation. An implication of these results is that common physiologic mechanisms may underlie spontaneous activity as imaged with fMRI in humans and slowly propagated activity as studied in animals.
© 2015 MedicalXpress.com