Targeting mutant proteins might be silver bullet for neurodegenerative diseases

Scientists have unraveled how mutant molecules damage the nervous system of people with Charcot-Marie-Tooth (CMT) disease, a group of disorders that hinder people's ability to move and feel sensation in their hands and feet, according to a paper published October 21, 2015 in Nature.
In laboratory testing, the researchers were able to improve symptoms of the disease in mice, raising hopes that they may have found an avenue for treating people with CMT.
The research, a collaboration between scientists from the Salk Institute and The Scripps Research Institute (TSRI), offers promising targets for developing new drugs for the disease. The findings may offer clues to understanding and treating other neurodegenerative disorders, including other forms of CMT.
"This solves a long-running mystery of how a gene mutation damages the neurons that carry information from the spinal cord to our muscles, resulting in a range of sensory and movement problems," says Samuel Pfaff, a neuroscience professor at the Salk Institute and one of the senior authors on the paper with Xiang-Lei Yang, a professor at TSRI. "It's an exciting finding, as we were able in experiments to reverse the symptoms of the disease by targeting the activity of these proteins."
CMT is a group of hereditary disorders that affects about 1 in every 2,500 people in the United States, making it one of the most common inherited neurological diseases. While different forms of the disease vary in their symptoms and underlying genetic causes, the common thread is that CMT damages the nerves in a person's arms and legs.
Symptoms of the disease, which typically appear in adolescence or early adulthood, include muscle weakness and decreased muscle size, loss of sensation and deformities in the feet and legs. The symptoms typically first appear in the lower extremities, but eventually may move into the hands.
It was known from early work from the Yang lab that one form of the disorder was caused by a gene mutation that resulted in a misshapen form of an enzyme known as GlyRS that serves many roles in cells, but how this resulted in neural damage remained a mystery.
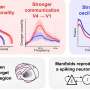
The Salk and TSRI scientists used a range of neuro-genetic, gene therapy, biochemical and structural biology research techniques to discover that the mutant GlyRS enzyme blocked molecular signals important for maintaining the health of motor neurons, the cells that carry messages from the brain to the muscles of the extremities. One surprise was that the GlyRS enzyme, which was commonly thought to remain inside neurons, was actually found outside the neurons. That is where it blocked a protein known as VEGF from connecting with a molecular sensor, named Nrp1, which is found on the surface of the neurons.
"Normally, these molecular messages essentially tell the neurons 'you're healthy,' stimulating them to remain robust and active," says Yang. "By blocking this message, the mutant GlyRS enzyme causes the motor neurons to go into decline and even die, which breaks the connection between the brain and the muscles in the limbs."
Having identified the mechanism by which the gene mutation leads to neuropathy, the researchers tried amplifying the "health" signal to see if they could override the blockade of the mutant GlyRS molecules. When they turned up the volume on the signal, using gene therapy techniques to increase the production of the signaling molecule, the mice with CMT regained muscle strength and their ability to walk improved significantly.
The findings suggest a possible avenue for developing new therapies for CMT by either dampening the activity of the mutant GlyRS enzymes or amplifying the competing signal that promotes the health of the motor neurons.
More broadly, the research suggests a new framework for exploring how the mutated proteins interfere with normal processes for neuron survival in other neurodegenerative diseases.
More information: CMT2D neuropathy is linked to the neomorphic binding activity of glycyl-tRNA synthetase, DOI: 10.1038/nature15510