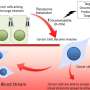
Single-cell analysis reveals subtypes of colorectal tumors
Combining single-cell genomics and computational techniques, a research team including Paul Robson, Ph.D., director of single-cell biology at The Jackson Laboratory (JAX), has defined cell-type composition of cancerous cells from 11 colorectal tumors, as well as adjacent noncancerous cells, a key to more targeted diagnosis and treatment.
"Using single-cell signatures," says JAX Research Scientist Elise Courtois, Ph.D., co-first author of a study published in Nature Genetics, "colorectal cancers can be further divided into subgroups based on cell-type composition of tumors. Because each of these subgroups has a different survival probability, our approach can provide oncologists with better information about prognosis and treatment options."
Advances in tumor genomic sequencing have improved classification of tumor subtypes, guiding more precise cancer treatments and improving patient survival. However, tumors typically contain a variety of cancerous and noncancerous cells that all contribute to the biology of the tumor.
To date, gene expression in such tumors has been profiled using bulk transcriptome methods, providing a single transcriptome measure for what, in essence, represents many cell types. By employing single-cell transcriptomic technology it is now possible to deconstruct a tumor into its component cell-type parts and therefore gain a better understanding of the underlying biology.
Robson and co-senior authors Shyam Prabhakar, a computational biologist at the Genome Institute of Singapore, and Iain Beehuat Tan, a medical oncologist at the National Cancer Centre Singapore, led an effort that screened 626 randomly selected individual cells from the colorectal tumors and adjacent normal cell samples using single-cell RNA sequencing.
Computationally examining each cell's transcriptome (the readout of all the messenger RNA molecules in that cell) using their new algorithm, the researchers identified two distinct subtypes of cancer-associated fibroblasts (CAFs). These CAFs significantly contributed to the mesenchymal gene expression found in bulk tumor transcriptome data, a signature more often associated with a cancer cell process known as epithelial-mesenchymal transition. Their data make the case that CAFs contribute to a worse prognosis in colorectal cancer patients.
These findings, Robson notes, show promise for even more refined classification of colorectal and other tumors in the future. "And as the cost of single-cell transcriptomic analysis continues to drop, oncologists can access better tumor profiling to guide the treatment of cancer patients," he says.
More information: Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors, Nature Genetics, nature.com/articles/doi:10.1038/ng.3818