Urinary tract infections reveal the importance of interactions between host susceptibility and bacterial gene expression

In the search for a defining feature of the Escherichia coli that cause urinary tract infections, comparative transcriptomics and infection models show that infection depends on both the host environment and gene expression levels in the bacteria.
Roughly 50 percent of women will suffer a urinary tract infection (UTI) at some point in their lives. For another 20 percent of those, the infection will recur at least once—and sometimes as often as every few months. UTIs are most often caused by strains of Escherichia coli bacteria known collectively as uropathogenic E. coli (UPEC), but the features enabling UPEC to infect the bladder are poorly defined.
To better understand UTIs and potentially offer new therapeutic strategies, a team of researchers analyzed 43 strains of E. coli sampled from 14 women with recurrent UTIs, and found that infection depends on both individual host susceptibility and how the bacteria express their genes. This new view on infection may also be applicable to other pathogens that lack a defining genetic signature. The study, published in Science Translational Medicine, was led by Henry Schreiber IV, a graduate student from Washington University and visiting scientist at Broad Institute, Jonathan Livny, senior research scientist in the Infectious Disease and Microbiome Program, Ashlee Earl, group leader for the Bacterial Genomics Group, and Scott Hultgren of Washington University.
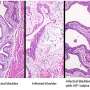
To examine how well the sampled UPEC could infect hosts in a controlled setting, the researchers tested the virulence of 21 strains in lab mice. The team saw a range of infection severity—but there were some surprising results.
Previous studies of UPEC have identified genes believed to give E. coli a potent ability to colonize the bladder. However, many E. coli strains that severely infected the mouse models were missing these virulence genes, and some of these strains outcompeted those with high numbers of the genes. The findings suggest that the role of these urovirulence factors may be important in only one of multiple strategies used by E. coli to infect the bladder.
Further comparative genomic and transcriptomic analyses showed infection severity correlated with the expression levels of certain genes shared by nearly all E. coli, including genes involved in motility and nutrient utilization. According to the researchers, sequence variations that affect gene expression, rather than protein-coding potential, appear to be key drivers of urovirulence.
Additionally, when tested in a second mouse model, some of the UPEC strains that successfully infected the first caused a less severe outcome. Based on these findings, the research team proposed a new pathogenesis model in which each combination of host and UPEC strain is a unique pairing of bacterial urovirulence potential and host susceptibility—and the compatibility of that pairing ultimately determines UTI outcome, including severity.
Thus, two women with UTIs could have infections caused by similar or radically different bacteria, and an E. coli strain that causes a UTI in one person may not make a different person sick at all due to differences in genetics, behavior, medical history, or other environmental features. This integrated analytical approach sets the stage for future studies using multiple angles of inquiry to better understand the complex dance between host and pathogen.
More information: Henry L. Schreiber et al. Bacterial virulence phenotypes ofand host susceptibility determine risk for urinary tract infections, Science Translational Medicine (2017). DOI: 10.1126/scitranslmed.aaf1283