New model can predict drug interactions and side effects even between a large number of components

Drug cocktails such as those for treating cancer, like the alcoholic versions offered at the local bar, are best when the proper ingredients are mixed in the right proportions. And like the cocktails we normally drink, the combination of ingredients can be better than the sum of its parts—or it can leave us with unwanted side effects. A new model developed in the group of Prof. Uri Alon of the Weizmann Institute of Science's Molecular Cell Biology Department can simplify the process of identifying the optimal blends for drug cocktails - even when a large number of ingredients is called for.
Drug cocktails - both antibiotic and anti-cancer - are increasingly used, among other things, because simultaneously attacking pathogenic cells with several different methods can reduce the risk of drug resistance. And doctors and pharmaceutical companies are interested in the advance of drug "mixology" because it can help create novel applications for existing drugs, since new ones are costly to develop and slow to reach the market.
But adding drugs together does not generally result just in the sum of their effects. For instance, one drug can alert mechanisms in a cell that pump the other drugs out of the cell, thus changing the dose at which the other drugs will be effective. Conversely, side effects can add up, so researchers often want to identify the lowest possible dose of any given drug. And with typically four or more drugs added together in chemotherapy cocktails, the number of possible combinations and doses is astronomical: It would be impossible to test them all to arrive at the optimal mix. This hurdle is known as the combinatorial explosion problem.
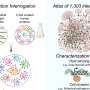
Because of the combinatorial explosion problem, say research students Anat Zimmer and Itay Katzir, who led the study, drug cocktails are often concocted without any good way of predicting the end result. Zimmer, Katzir and Alon developed a method that bypasses the need for an astronomical number of measurements. Their method requires only a small number of measurements on drug pairs.
The tests are conducted on human cancer cells or bacteria grown in lab dishes. The group tested each drug - separately and in pairs - to understand the effects at several different doses. This enabled the researchers to determine how drug A affects the actions of B, and vice versa, and the new mathematical model the group developed was then extrapolated to predict the interactions among three, four and more drugs in combination. Further testing showed that the model performs better than existing methods of dealing with the combinatorial explosion problem. Thus researchers using the model would not need to test every possible combination to arrive at the optimal doses in drug cocktails.
"There is an urgent demand for methods that can predict how drug cocktails will work," says Katzir. "This model may take much of the expensive guesswork out of the process of developing such cocktails."
"The model might prove especially useful for personalized medicine - for example, in cancer -because each tumor can react differently to the same drugs," adds Zimmer. "It provides a way to mix that perfect cocktail without having to try out all of the possible combinations."
More information: Anat Zimmer et al. Prediction of multidimensional drug dose responses based on measurements of drug pairs, Proceedings of the National Academy of Sciences (2016). DOI: 10.1073/pnas.1606301113