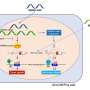
Household environment—not genetics—shapes salivary microbes
Researchers in the United Kingdom have discovered that the mix of microorganisms that inhabit a person's saliva are largely determined by the human host's household. The study, published this week in mBio, an open-access journal of the American Society for Microbiology, shows that early environmental influences play a far larger role than human genetics in shaping the salivary microbiome—the group of organisms that play a crucial role in oral and overall health.
"It's generally becoming known that there's a link between our microbiomes and our health and that's reason enough to find out what's in there, how they arrived there, and what they are doing," says Adam P. Roberts, senior lecturer in antimicrobial chemotherapy and resistance at the Liverpool School of Tropical Medicine. Roberts co-led the study, which was conducted during his previous post at the UCL Eastman Dental Institute. UCL Genetics Institute graduate student Liam Shaw adds, "The oral cavity is naturally colonized by hundreds of bacterial species, which stop external pathogens from establishing a foothold, but they also can themselves cause oral disease."
The research team wanted to know how the salivary microbiome gets established and which factors are most responsible for the mix of bacteria found there. Roberts' colleague, UCL immunologist Andrew M. Smith, had access to a unique sample set—DNA and saliva from an extended, Ashkenazi Jewish family living in various households spread across four cities on three continents. That allowed the team ask how much of the variation seen in salivary microbiomes is due to host genetics and how much is due to environment.
Because the family members are ultra-orthodox Ashkenazi Jews, they share cultural diets and lifestyles that control for many confounding factors. Also, because the family members' DNA had already been sequenced to the level of single changes in the DNA code, the research team had a unique and precise measurement of their genetic relatedness.
Next, Shaw and the team sequenced the bacterial DNA signatures present in saliva samples from 157 family members and 27 unrelated Ashkenazi Jewish controls. Across all samples, they found the core salivary microbiome made up of bacteria from the genera Streptococcus, Rothia, Neisseria, and Prevotella.
To get at what might be driving differences at the bacterial species level, Shaw and the team used statistical methods adopted from ecology to determine which factors are responsible for the most variation. When comparing factors such as shared household, city, age, and genetic relatedness, the factor that determined who shared the most similar saliva microbes was overwhelmingly household.
"What that tells us is that the contact and sharing of microbes that goes on at the very local environment is what determines the differences between individuals," says Shaw.
Spouses and parents and children younger than 10 living in a household together had the most similar saliva microbiomes. "The contact doesn't even have to be intimate, like kissing," says Roberts. "Individuals' hands are covered in saliva and they are touching everything in the house." Children younger than 10 had more similar bacteria to their parents than older children, perhaps reflecting that older children are becoming "more independent individuals," says Roberts.
The team also looked carefully at whether genetic relatedness drove the makeup of the saliva microbiome. When they used a measure of relatedness based on family tree relationships alone, they saw a small, but statistically significant effect. However, when they used the genetic sequence information, a more accurate measure of relatedness, the effect disappeared. In other words, a person's genetics played virtually no role in shaping their saliva microbes.
"Pedigrees do not always precisely reflect actual genetic relatedness in terms of the amount of genome shared," says Shaw. Roberts encourages other researchers undertaking large-scale microbiome studies to use detailed human genetic sequence information rather than relying on family trees.
This study shows that environments shared during upbringing play a major role in determining what community of bacteria gets established. And knowing that the shared environment drives the microbiome, Roberts says, may gives us the ability to one day modulate it.
He points to the example of periodontitis, or gum disease, an incredibly common and often debilitating infectious disease associated with an altered microbiome. "Once we understand the members of the microbiome that are responsible for health, our everyday behavior could change to shift our microbiome favorably."