'Mix and match' assembly instructions guide immune cell attacks

Institute researchers have discovered how immune cells use a unique set of assembly instructions to 'mix and match' how they respond to, and kill, tumour and diseased cells.
Cell surface receptor complexes are like Lego pieces, constructed using different molecular combinations to guide how the immune cell acts when it makes contact with a cancerous cell, an infection, or other external signals. The new research identifies the features that allow these pieces to assemble in specific combinations.
Understanding how these complexes assemble 'naturally' could pave the way for future improvements in immunotherapy, such as engineering cancer-specific immune killers.
Associate Professor Matthew Call and Dr Melissa Call from the Walter and Eliza Hall Institute led the research with colleagues in Spain and the US, published in the journal Proceedings of the National Academy of Sciences.
Associate Professor Matthew Call said the team discovered an entirely new set of assembly instructions used by molecular sensors embedded in the thin fatty cell membrane to build receptor complexes in response to different stimuli.
"Effectively, these membrane-bound sensors determine 'who' the immune cell talks to. This is really important in the promising field of cancer immunotherapy, because it could help us better engineer cells to specifically talk to – and destroy – cancer cells," Associate Professor Matthew Call said.
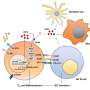
Cell surface receptor complexes consist of an external receptor that binds to signalling molecules, an internal molecule that instructs the cell how to respond, and a cell membrane-embedded portion that anchors and links the other two segments.
In the past, these cell membrane-embedded sections of the receptor were largely ignored, partly because they are so difficult to work with, Associate Professor Matthew Call said.
"However more and more we are discovering what a critical role they play in how signals are received and acted upon by the cell. In fact, the sensors found in the fatty cell membrane actually control how the cell responds to external signals by pulling different external and internal sections together like building blocks to drive the correct response," he said.
The researchers began by studying the receptors on natural killer (NK) cells, but found that the same assembly instructions were used in a host of immune cells that "run around and eat and blow up" cancerous and other diseased cells, Dr Melissa Call said.
"One subset of these receptors, called Fc receptors, were the focus of this research. We were particularly looking at the subset of Fc receptors found on natural killer (NK) cells – immune cells that poison tumour and virus-infected cells that have been 'marked' by antibodies," Dr Melissa Call said.
She said the study showed that different subsets of Fc receptors used completely different assembly instructions compared to other, similar receptors.
"Over the past decade or so, this has become really important therapeutically because of a new field of cancer immunotherapy called chimeric antigenic receptor therapy, or CART," Dr Melissa Call said.
"The idea of CART is that you create specially engineered receptors in immune cells that are highly specific for an individual cancer. Understanding in depth how these receptors are assembled naturally is vital for us to understand how best to design them ourselves for cancer therapy, to look at improved ways of stimulating the immune response to cancer."
Dr Melissa Call said the laboratory's long-term goal was to decipher and control how immune receptors work, to find new ways of manipulating the immune response for therapeutic treatments.
"Controlling how the immune receptors are assembled could allow us to boost the immune response in the case of infections or immune deficiencies, dampen it down in the case of autoimmune disease, or redirect the immune response to better target cancers," she said.
More information: Alfonso Blázquez-Moreno et al. Transmembrane features governing Fc receptor CD16A assembly with CD16A signaling adaptor molecules, Proceedings of the National Academy of Sciences (2017). DOI: 10.1073/pnas.1706483114