December 12, 2017 report
Association found between abnormal cerebral connectivity and variability in the PPARG gene in developing preterm infants

(Medical Xpress)—A team of researchers with King's College London and the National Institute for Health Research Biomedical Research Centre, both in the U.K., has found what they describe as a strong association between abnormal cerebral connectivity and the PPARG gene and uncharacteristic white matter development in the brains of preterm infants. In their paper published in Proceedings of the National Academy of Sciences, the group describes their use of unbiased learning analysis on data obtained from 272 preterm infants and what they learned from it.
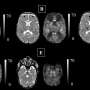
Babies born early tend to have more medical issues than those that make it to full term, and the earlier they come, the more hurdles they face. For babies born at 33 weeks or less, there is also an increased risk of abnormal brain development. As the researchers note, a lot of research regarding abnormal brain development in preterm infants has been done, but still little is known about the process itself. What is known is that it is usually related to an excess amount of white matter (tissue in the brain made up mostly of nerve fibers and surrounded by myelin) developing in such infants. In this new effort, the researchers took a new approach to better understanding the root cause of the abnormality.
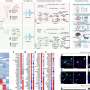
The new approach involved accessing medical data on 272 infants born at 33 weeks or less, and who had undergone both MRI scans and genomic testing. The goal of the effort was to find out if there was a genetic element involved in the abnormal brain development, and if so, to locate the genes responsible. The work by the team involved combining data from the genetic analysis with the imaging data, which the team referred to as an unbiased machine learning approach. The idea was to find out if there was a genetic factor common to all of the infants with abnormal brain development.
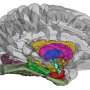
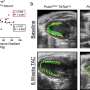
The analysis revealed a strong connection between cerebral connectivity and differences in the PPARG gene, which, the researchers note, implicated PPARG signaling as the reason for the development of excess white matter. This finding means that it might be possible to develop drugs that target PPARG in preterm infants, preventing the development of excess white matter and hopefully, neurocognitive problems.
More information: Michelle L. Krishnan et al. Machine learning shows association between genetic variability inPPARGand cerebral connectivity in preterm infants, Proceedings of the National Academy of Sciences (2017). DOI: 10.1073/pnas.1704907114
Abstract
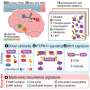
Preterm infants show abnormal structural and functional brain development, and have a high risk of long-term neurocognitive problems. The molecular and cellular mechanisms involved are poorly understood, but novel methods now make it possible to address them by examining the relationship between common genetic variability and brain endophenotype. We addressed the hypothesis that variability in the Peroxisome Proliferator Activated Receptor (PPAR) pathway would be related to brain development. We employed machine learning in an unsupervised, unbiased, combined analysis of whole-brain diffusion tractography together with genomewide, single-nucleotide polymorphism (SNP)-based genotypes from a cohort of 272 preterm infants, using Sparse Reduced Rank Regression (sRRR) and correcting for ethnicity and age at birth and imaging. Empirical selection frequencies for SNPs associated with cerebral connectivity ranged from 0.663 to zero, with multiple highly selected SNPs mapping to genes for PPARG (six SNPs), ITGA6 (four SNPs), and FXR1 (two SNPs). SNPs in PPARG were significantly overrepresented (ranked 7–11 and 67 of 556,000 SNPs; P < 2.2 × 10−7), and were mostly in introns or regulatory regions with predicted effects including protein coding and nonsense-mediated decay. Edge-centric graph-theoretic analysis showed that highly selected white-matter tracts were consistent across the group and important for information transfer (P < 2.2 × 10−17); they most often connected to the insula (P < 6 × 10−17). These results suggest that the inhibited brain development seen in humans exposed to the stress of a premature extrauterine environment is modulated by genetic factors, and that PPARG signaling has a previously unrecognized role in cerebral development.
© 2017 Medical Xpress