Protein associated with ALS points to possible targets for therapeutic intervention

Scientists at the University of Alberta may have found possible targets for therapeutic interventions in the fight against Lou Gehrig's disease.
Biophysicist Michael Woodside and his research team conducted the first single-molecule study of folding in the protein superoxide dismutase-1 (SOD1), an antioxidant whose misfolding is linked to the neurodegenerative disease ALS. They found that it has much more complex folding than previously thought.
The results suggest an explanation for the protein's propensity to misfold similarly to proteins in prion diseases.
"When we pulled the protein apart, we were expecting its structure to come apart all at once based on what was previously known, but what we found instead was a mess," said Woodside, professor in the U of A's Department of Physics. "But clearer patterns started to emerge after unfolding and refolding it several thousand times. You get a lot more detail working at this single-molecule scale, and it allows us to start piecing the whole picture together."
Best known for his work on prion diseases such as mad cow disease and the associated human form of Creutzfeldt-Jakob, Woodside said he and his colleagues were drawn to the problem based on the protein's prion-like characteristics, noting that the behaviour in the misfolding is reminiscent of mad cow disease.
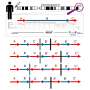
Woodside used similar techniques from his previous work to better understand SOD1, using laser tweezers to measure the unfolding and refolding of single molecules.
He explained that there was a stable core of the protein that was the last to unfold and the first to refold.
"What we are finding is that when it folds up to an incorrect state, it actually always starts off making the same stable core that you find when it goes into the correct state. It just takes a wrong turn partway down that pathway," he noted.
It was in the misfolding around this core where he and his team were trying to pinpoint the wrong turns of the folding pathways of the protein. They identified several types of misfolded pathways and resolved numerous previously undetected intermediate states en route to a more complete map.
"When you don't understand why something is misfolding, it becomes difficult to target therapeutic treatments. So understanding where things are going wrong helps the targeting process become more rational rather than leaning on random screening," said Woodside.
Woodside said the next challenges are to scale up the findings, connecting single molecules to an entire cell and then finally to a full organism. He is now working with a leading ALS clinician out of the University of British Columbia to advance future work, focusing on how mutations that lead to inherited forms of ALS can cause misfolding to spread from molecule to molecule and cell to cell.
"Partially native intermediates mediate misfolding of SOD1 in single-molecule folding trajectories" was published in the Dec. 1 issue of Nature Communications.
More information: Supratik Sen Mojumdar et al, Partially native intermediates mediate misfolding of SOD1 in single-molecule folding trajectories, Nature Communications (2017). DOI: 10.1038/s41467-017-01996-1