New study links genetic diversity of tumors with resistance to treatment in Asian lung cancer patients
Scientists from A*STAR's Genome Institute of Singapore (GIS) and medical oncologists from the National Cancer Centre Singapore (NCCS) have discovered that lung cancer tumours in Asian patients contain much higher genetic diversity than previously expected. Hence, this type of tumour tends to develop resistance despite initial tumour shrinkage. With this discovery, scientists and oncologists are better equipped to guide treatments and develop more refined and personalised approaches to common non-small-cell lung carcinoma (NSCLC). The study was published in the international journal Nature Communications.
Lung cancer is the most common cause of cancer mortality in the world. It accounts for approximately 19 percent of all cancer-related deaths worldwide and has a mortality rate more than twice that of any other cancer. Lung cancer is also the most common type of cancer in Asia2. The number of incidences and deaths are expected to rise with increased pollution, particularly in large, densely populated Asian cities. Thus, newer and more effective individualised treatment strategies are crucial to tackle the growing threat and improve the quality of patients' lives.
Lung cancer in Asian patients is characterised by an important mutation in a gatekeeper gene known as the epidermal growth factor receptor, EGFR3. Gatekeeper genes, also known as the tumour suppressor genes, control and regulate cell cycles. It was discovered that mutations in this gene occur in more than 50 percent of the tumours in Singaporean lung cancer patients. While drugs targeting it are effective in controlling the disease, the response is short-lived. Most patients eventually succumb to cancer relapse in a matter of months or a few years. In some instances, patients do not even respond to these drugs at all. The variability in clinical outcomes, and the tumours' seemingly inevitable development of resistance to treatment, have intrigued doctors for years, and have remained major stumbling blocks in administering better patient care.
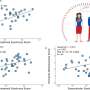
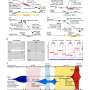
"This joint study is one of the first major efforts to characterise and identify lung tumours in Singaporean patients on a large scale. It has generated a treasure trove of new genetic information and enabled us to perform detailed analyses, leading us to conclude that lung tumours in Asian patients are surprisingly more complex than previously appreciated," mentioned Dr. Rahul Nahar, the first author of this study and a Research Associate at GIS.
"The study of the genetic complexity of tumours in Asian patients has provided us with new insights as to why they may quickly develop resistance after initial response to anti-EGFR drug inhibitors. We also found that tumours with a high number of drivers, mutations that cause cancer progression, tend to be associated with poor drug response," explained Dr. Axel Hillmer, Principal Investigator at GIS and a co-corresponding author of this study.
"Understanding the comprehensive genetic landscape of these tumours allows us to go beyond single gene mutations (such as EGFR) to better understand the behaviour of individual tumours, and tailor treatments more effectively. Further work needs to be focused on identifying drug combinations or treatment strategies that take into account the tumours' ability to adapt to different treatments," said Dr. Daniel Tan, Senior Consultant Medical Oncologist at NCCS and the corresponding author of this paper.
Professor Ng Huck Hui, Executive Director of GIS, said, "Timely and comprehensive research on disease states is critical, especially for diseases with a mortality rate as high as lung cancer. The study represents one of the first attempts at such an in-depth analysis of this unique subtype of NSCLC, and has yielded new insights and research directions. Discoveries like this will continue to pave the way for developing increasingly precise treatments."
More information: Rahul Nahar et al. Elucidating the genomic architecture of Asian EGFR-mutant lung adenocarcinoma through multi-region exome sequencing, Nature Communications (2018). DOI: 10.1038/s41467-017-02584-z