A non-coding RNA lasso catches proteins in breast cancer cells
A Danish-German study reports new findings about long non-coding RNA expression in the development of cancer. The results have an impact on the understanding of dynamic regulation of gene expression in biological processes.
Cells are divided into several compartments with specialized functions. DNA contains genetic information, and is localized in the cell nucleus where gene production is controlled. The nucleus is divided into the soluble nucleoplasm and the insoluble chromatin. Messenger RNA (mRNA) encodes for proteins, and the process of translation occurs in the cytoplasm. Long non-coding RNAs primarily work in the nucleus, where they are essential components of controlling gene expression.
The research team has paid particular attention to long non-coding RNAs that can enhance the production of specific mRNAs, and hence proteins, in breast cancer cells. They show that the expression of long non-coding RNAs can result in particularly high expression levels of specific proteins with involvement in cancer.
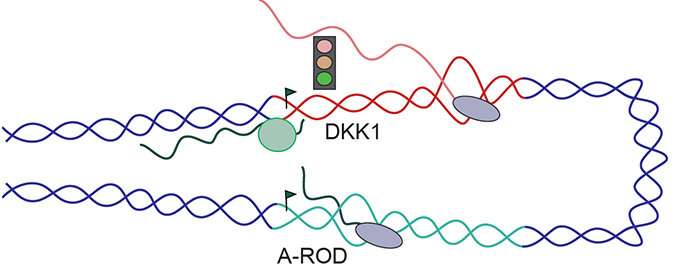
The researchers report that in the development of cancer, it matters where and how much of a particular long non-coding RNA is produced, but also how the long non-coding RNA is dynamically distributed within the cell. The authors show that the long non-coding RNA called A-ROD (Activating Regulator Of DKK1 expression) is only functional the moment it is released from chromatin into the nucleoplasm. At this transient phase, it can bring transcription factors—proteins controlling the synthesis of other genes—to specific sites in DNA to enhance gene expression. After its complete release from chromatin, A-ROD is no longer active as an enhancing, long non-coding RNA. In a way, A-ROD functions as a lasso that can be thrown from DNA to catch proteins.
The results are exciting both from an experimental and therapeutic point of view, as the strategies for targeting RNA expression in the cytoplasm, the nucleoplasm and at the chromatin differ widely. The team behind the study believes that these differences can be exploited to optimize the approaches for targeting RNA-dependent processes in disease. A future scientific goal is to identify more of the non-coding RNA lassoes to fully understand their potential and application in regulation of gene expression.