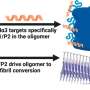
The role of 'extra' DNA in cancer evolution and therapy resistance
Glioblastoma (GBM) is the most common and aggressive form of brain cancer. Response to standard-of-care treatment is poor, with a two-year survival rate of only 15 percent. Research is beginning to provide a better understanding of the processes underlying cell-to-cell differences within GBM tumors - a crucial finding because these differences contribute to therapy resistance.
The ultimate goal is to identify what pathways can be targeted to block glioma progression.
To this end, a multi-institutional team led by Professor Roel Verhaak, Ph.D., of The Jackson Laboratory (JAX) and Ana C. deCarvalho, Ph.D., Assistant Professor from Henry Ford Health System's Hermelin Brain Tumor Center in Detroit, MI, tracked genomic alterations detected in patient samples during tumor cell evolution in culture, in patient-derived xenograft (PDX) mouse models from the cultures, as well as before and after treatment in patients. In a recent paper in Nature Genetics, the team reports that tumor progression was often driven by cancer-promoting genes, known as oncogenes, on extrachromosomal pieces of DNA.
Detailed analyses of the tumor cells from patient to culture to mouse revealed that, for the most part, the cells retained the same genomic lesions. This is good news overall, as it indicates that PDX mice can provide a relatively accurate and effective experimental platform for GBM. The primary caveat was the finding that in a few cases, the numbers of oncogene copies differed between tumors and the cultures and PDX mouse samples derived from them. If an oncogene is increased, or amplified, that can both cause and maintain cancer, so differences in gene amplification can be very important.
But why did the levels of oncogene amplification change? What the researchers found was that the differences were caused by oncogenes that weren't part of chromosome sequences as usual. Instead, they were on separate circular pieces of DNA, known as extrachromosomal (ec) DNA. These pieces of DNA are not found in normal cells, and cause major increases in the expression of oncogenes. More detailed investigation showed that many instances of oncogene amplification found in the glioma tumors involved ecDNA elements.
"The selective advantage conferred to tumor cells by the regulation of oncogene copy number in ecDNA has not been sufficiently addressed in interpreting results in the laboratory or in clinical trials. Using the GBM patient-derived models carrying ecDNA amplification of the most frequent oncogenes, we are developing and testing novel combination therapies specific for each unique tumor," says deCarvalho.
ecDNA elements were first observed directly under microscopes in cancer cells more than 50 years ago, but it remains unknown how they arise in the first place. Technological limitations have impeded studies of ecDNA in detail, despite a recent publication in Nature suggesting their presence in nearly half of cancers. In fact, their role in disease has not been extensively studied, but it's an important topic. Unlike chromosomal DNA, ecDNA is inherited inconsistently as a tumor grows. That is, when a cancer cell divides, the DNA on the chromosomes almost always gets accurately duplicated and remains the same in the daughter cells. But ecDNA inheritance appears to be far more random. Sometimes both daughter cells inherit ecDNA, but sometimes all or most of it will end up in one cell and not the other.
"The process quickly creates important differences between cells within the same tumor, and it helps accelerate the evolution of the cancer," says Verhaak. "It provides the cells with more ways to evade stress. Therefore, there's a better chance that at least some of the cells will survive severe stress, such as stresses caused by a chemotherapy or radiation."
One reason ecDNA has been relatively ignored is that it's hard to detect using standard sequencing methods, which don't accurately detect and separate it from chromosomal DNA. But it's now attracting more attention, and the work moving forward will likely help explain why cancers such as GBM are difficult to treat and evolve therapy resistance so rapidly.
"We think targeting ecDNA has huge potential for the development of new cancer treatments" says Verhaak. "We're now working to develop sequencing-based protocols to identify ecDNA more efficiently. The bigger goal is to learn how and why ecDNA elements form. If we can block those mechanisms, we'll have a way to prevent the evolution, and perhaps even the formation, of many cancers."
More information: Ana C. deCarvalho et al, Discordant inheritance of chromosomal and extrachromosomal DNA elements contributes to dynamic disease evolution in glioblastoma, Nature Genetics (2018). DOI: 10.1038/s41588-018-0105-0