Two thousand human brains yield clues to how genes raise risk for mental illnesses
It's one thing to detect sites in the genome associated with mental disorders; it's quite another to discover the biological mechanisms by which these changes in DNA work in the human brain to boost risk. In their first concerted effort to tackle the latter, 15 collaborating research teams of the National Institutes of Health-funded PsychENCODE Consortium leveraged statistical power gained from a large sample of about 2000 postmortem human brains.
The teams published their findings in seven research articles, spotlighted on the cover of a "psychiatric genomics" special issue of Science—along with two in Translational Medicine and one in Science Advances—on Dec. 14, 2018. In addition, the consortium is sharing their data with the research community via the online PsychENCODE Knowledge Portal .
Applying newly uncovered secrets of the brain's molecular architecture, they developed an artificial intelligence model that's six times better than previous ones at predicting risk for mental disorders. They also pinpointed several hundred previously unknown risk genes for mental illnesses and linked many known risk variants to specific genes.
"For the first time, we now have a beginning of an understanding of the biology, the molecular pathophysiology, of schizophrenia, bipolar disorder and autism spectrum disorder (ASD)," said Thomas Lehner, Ph.D., M.P.H., director of the Office of Genomics Research Coordination (OGRC) of the National Institute of Mental Health (NIMH), which launched the PsychENCODE initiative in 2015.
In brain tissue and in single cells, the researchers examined patterns of gene expression (transcriptome), marks of gene regulation (epigenome), as well as genetic variants linked to mental illnesses in genome-wide association studies.
"The consortium's integrative genomic analyses elucidate the mechanisms by which cellular diversity and patterns of gene expression change throughout development and reveal how neuropsychiatric risk genes are concentrated into distinct co-expression modules and cell types," explained Nenad Sestan, M.D., Ph.D. , of Yale University, New Haven, Connecticut, leader of one of the teams, in an accompanying editorial in Science.
The implicated variants are mostly small-effect genetic variations that fall within regions of the genome that don't code for proteins, but instead are thought to regulate gene expression and other aspects of gene function. PsychENCODE sought to uncover the specific roles played by these elusive actors at particular time points and places in the developing brain.
The researchers examined brain tissue and molecules from prenatal development as well as from people with schizophrenia, bipolar disorder, ASD and typical development – and compared findings with parallel data from non-human primates. They also incorporated data from related NIH initiatives in humans, including ENCODE and GTEx, which enhanced the statistical power of the analysis.
Among the key findings:
- Gene variants linked to mental illnesses exert more effects when they collectively form "modules"—groups of co-expressed, communicating genes with related functions—in specific cell types and brain areas, and at specific developmental time points that seem to coincide with the course of illness. For example, ASD-linked modules were seen early in prenatal development—possibly related to the early onset of symptoms—while schizophrenia-linked modules formed later, plausibly accounting for the onset of symptoms in late adolescence or early adulthood.
- One suspect module (ME37) in particular showed rapid change in a late prenatal "transition" period. It harbored genes and transcription factors linked to multiple neurodevelopmental disorders, including ASD and schizophrenia, to traits such as neuroticism and IQ—and to key processes, such as the birth of new neurons and epigenetic regulation of gene expression and behaviors including learning and memory
- Variability in risk gene expression and cell types peaked during formative stages in early prenatal development and again during the teen years—a pattern also seen in non-human primates.
- Mental illness risk genes are among genes found to be unique to humans.
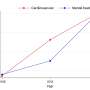
- An artificial intelligence computational learning model that integrated postmortem brain-derived data increased researchers' ability to retrospectively predict a person's risk for developing a mental illness to about 25 percent (better than chance), compared to only about 4 percent with previous models based on genetic data alone. This "multi-omic" model integrates postmortem data across the transcriptome, epigenome, and proteome—in addition to genome data.
- A module with many genes coding for brain immune cells showed a pattern of dysregulated gene expression linked to mental illnesses—excess expression in ASD and weak expression in schizophrenia and bipolar disorder. This, and other signs, added to evidence linking the disorders to inflammation in the brain.
- In postmortem brains of people with a mental illness, thousands of RNAs, which are molecules of gene expression, were found to have anomalies.
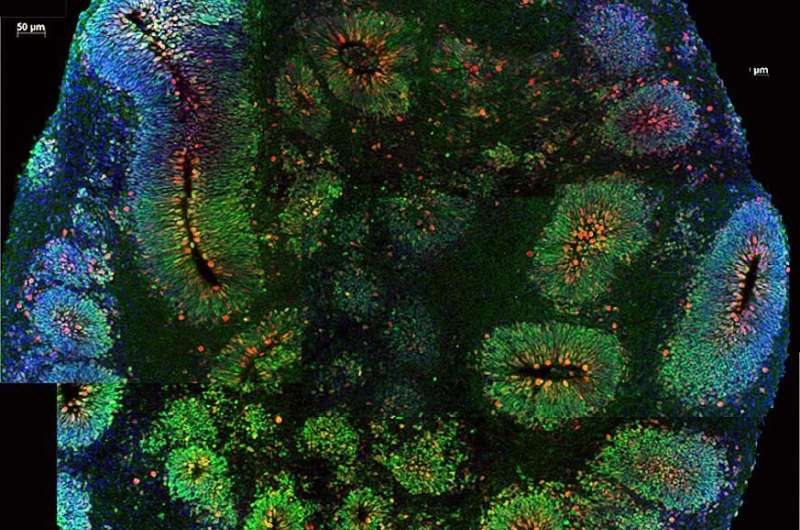
- ASD risk gene variants and the regulators they disrupt were highly expressed in organoids, brain-like tissue from human cells growing in a dish, that mimic the cortex during early prenatal development, when modules linked to ASD are active.
"The multi-omic data resource generated by the PsychENCODE collaboration will pave a path for building molecular models of disease and developmental processes and may provide a platform for target identification for pharmaceutical research," added the NIMH OGRC's Geetha Senthil, Ph.D., who served as program officer for the first phase of the project.
More information: Qingtuan Meng et al. The DGCR5 long noncoding RNA may regulate expression of several schizophrenia-related genes, Science Translational Medicine (2018). DOI: 10.1126/scitranslmed.aat6912
Chao Chen et al. The transcription factor POU3F2 regulates a gene coexpression network in brain tissue from patients with psychiatric disorders, Science Translational Medicine (2018). DOI: 10.1126/scitranslmed.aat8178
Suhn K. Rhie et al. Using 3D epigenomic maps of primary olfactory neuronal cells from living individuals to understand gene regulation, Science Advances (2018). DOI: 10.1126/sciadv.aav8550
Joon-Yong An et al. Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder, Science (2018). DOI: 10.1126/science.aat6576
Mingfeng Li et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks, Science (2018). DOI: 10.1126/science.aat7615
Michael J. Gandal et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder, Science (2018). DOI: 10.1126/science.aat8127 Daifeng Wang et al. Comprehensive functional genomic resource and integrative model for the human brain, Science (2018). DOI: 10.1126/science.aat8464
Ying Zhu et al. Spatiotemporal transcriptomic divergence across human and macaque brain development, Science (2018). DOI: 10.1126/science.aat8077
Anahita Amiri et al. Transcriptome and epigenome landscape of human cortical development modeled in organoids, Science (2018). DOI: 10.1126/science.aat6720
Prashanth Rajarajan et al. Neuron-specific signatures in the chromosomal connectome associated with schizophrenia risk, Science (2018). DOI: 10.1126/science.aat4311
Joon-Yong An et al. Genome-wide de novo risk score implicates promoter variation in autism spectrum disorder, Science (2018). DOI: 10.1126/science.aat6576