Mapping genetic variants driving toxicity to leukemia therapy

Scientists at St. Jude Children's Research Hospital, alongside collaborators around the world, have created a comprehensive reference of functional variants in an important drug-metabolizing enzyme called NUDT15. This thorough understanding of NUDT15 variants provides an invaluable resource for predicting which patients being treated with thiopurine drugs for acute lymphoblastic leukemia (ALL) are likely to experience toxicity. The work was published today in Proceedings of the National Academy of Science.
Thiopurines are important antimetabolite drugs, which make up a critical part of curative treatment for ALL. Genetic variations can directly influence drug toxicity and effectiveness of thiopurines. Researchers at St. Jude and elsewhere previously identified NUDT15 variants as a major genetic cause of toxicity during thiopurine therapy, particularly in people of Asian and Hispanic descent.
Pharmacogenetics is a field of science that uses genomic data to guide precision medicine. To implement genetic-guided treatment, researchers need a rigorous and thorough understanding of all the variants and their functions. The vast majority of genetic variants in the human genome remain uncharacterized. These variants are considered of unknown significance, leaving a void in our knowledge about how to implement precision medicine, including pharmacogenetics, for these patients.
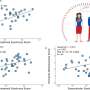
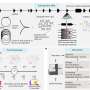
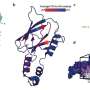
"To address this issue, we designed a high throughput experimental system to characterize the functional consequences of 91% of all possible genetic variants of the NUDT15 gene," said senior author Jun J. Yang, Ph.D., of the St. Jude Departments of Pharmaceutical Sciences and Oncology. "In the span of only a few months, we experimentally studied close to 3,000 variants in this gene in parallel and found that 30% are damaging, which we predicted to cause thiopurine toxicity in patients."
Variants identified in this way (by function) more accurately predict toxicity in patients treated with thiopurines than bioinformatic algorithms can. Using patient data with a subset of these variants, the researchers found that NUDT15 variants predicted by this method to cause toxicity, did induce such toxicity. Conversely, the variants predicted to have benign effects did not cause toxicity. While the authors caution that future studies of larger sample sizes are needed to validate these results, the initial findings show promise.
"This reference vastly improves the ability to implement pharmacogenetics-guided thiopurine treatment, not just for ALL but for other diseases treated with thiopurines such as inflammatory bowel disease," said first author Chase Suiter, formerly of St. Jude and now a graduate student at the University of Washington. "Importantly, the scale of this type of variant-scanning experiment was almost unimaginable just a couple of years ago, and we are excited about applying it to study many other genes related to drug response."
More information: Chase C. Suiter et al. Massively parallel variant characterization identifies NUDT15 alleles associated with thiopurine toxicity, Proceedings of the National Academy of Sciences (2020). DOI: 10.1073/pnas.1915680117