Predicting immunotherapy success
One of the frustrations with anti-cancer therapy is that no one drug fits all: Most work well in some people but have little effect in other patients with the same type of cancer. This is as true of the newer immunotherapy treatments as it is of older types of chemotherapy. Weizmann Institute of Science researchers have now identified new markers that can help predict which patients have a better chance for a positive response to immunotherapy treatments. Their findings were reported in Nature Communications.
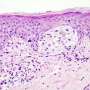
For Prof. Yardena Samuels and her research group partnered with Prof. Eytan Ruppin's lab in the National Institutes of Health (USA), basing their research on the understanding that learning to predict which treatments are most likely to work is the first step to creating a personalized approach to curing cancer. The focus of their work is melanoma—a collection of skin cancers that are often hard to treat, and which may be made up of varied tumor cells containing hundreds of different mutations. A certain percentage of melanomas have in recent years been successfully treated with immunotherapy drugs known as checkpoint inhibitors, which work by removing internal "obstacles" that trip up the body's own immune system and keep it from attacking the cancer. Unfortunately, for others, these drugs remain ineffective.
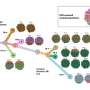
To understand the differences in response between different people, the research team, led by postdoctoral fellow Dr. Shelly Kalaora in Samuels' group and Joo Sang Li in the Ruppin group, first analyzed data from 470 melanoma patients that has been made available in the Cancer Genome Atlas (TCGA). They were particularly interested in differences in survival rates of patients in whom particular subunits of "immunoproteasomes" are overexpressed compared to those with low expression of these subunits. These are a variation on the proteasome subunits normally expressed in most cells (except for immune cells). Proteasomes are protein complexes that function as "cutters," reducing long proteins to short pieces called peptides. These peptides are later presented on the cells' surface by molecules called human leucocyte antigens (HLA). HLA peptides are basically bits of "information"—small molecules displayed on the outsides of cells—that "report" about new threats that the immune system needs to assess and address.
The immunoproteasome is assembled out of altered subunits and thus produces a unique collection of HLA peptides. The team thought that a particular change and overexpression in the HLA peptide repertoire might result in better recognition of the tumor cells by the immune system and thus better elimination of cancer cells.
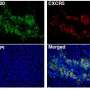
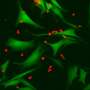
To test this idea, the researchers cultured lines of tumor cells from melanoma patients in which they overexpressed the immunoproteasome subunits and identified the HLA peptides presented in each situation. By testing the response of the immune cells from the same patients, they showed that the newly formed HLA peptides were indeed more reactive compared to the HLA peptides presented by cells without this overexpression.
Could two subunits the research identified in particular and the unique HLA-bound peptides these produce be a predictor of immunotherapy success? The experiments showed that in tumor cells in which the subunits were overexpressed, the various immune system components that directly fight the cancer were more prevalent and more active than average. Indeed, looking back at the details of cancer patients in the database, the team reported that the expression levels of the two subunits were excellent predictors of the outcome—better than the tumor mutational burden, a biomarker that is currently used in the clinic. The researchers in Samuels's and Ruppin's lab suggest that expression of the immunoproteasome may be used as a biomarker for predicting better outcomes in melanoma and, together with mutational load testing, may improve patient matching to currently available immunotherapy.
More information: Shelly Kalaora et al. Immunoproteasome expression is associated with better prognosis and response to checkpoint therapies in melanoma, Nature Communications (2020). DOI: 10.1038/s41467-020-14639-9