Epstein-Barr virus rewires host epigenomes to drive stomach cancer
The Epstein-Barr virus (EBV), one of the most common human viruses, is associated with about 8 to 10% of stomach—or gastric—cancers, the third leading cause of cancer death globally. Researchers from Chiba University in Japan, Duke-NUS Medical School, Singapore, and the Agency for Science, Technology and Research (A*STAR)'s Genome Institute of Singapore (GIS) have revealed a novel paradigm in EBV-associated gastric cancer, whereby the EBV viral genome directly alters the host epigenetic landscape to promote the activation of proto-oncogenes (genes involved in normal cell growth that can mutate into cancer-causing genes) and tumorigenesis.
The human genome is the complete set of human genetic information, and the epigenome describes modifications to the genome that determine whether genes are turned on or off when and where they are needed. Unlike genetic information, the epigenome is dynamic and responsive to external stimuli; certain external stimuli can cause abnormal DNA modifications which, in turn, can disrupt normal gene expression and contribute to cancer development.
The research group, led by senior and co-corresponding authors, Dr. Atsushi Kaneda, Professor at the Graduate School of Medicine, Chiba University, and Dr. Patrick Tan, Professor at the Program in Cancer and Stem Cell Biology, Duke-NUS Medical School, and Executive Director of GIS, conducted a comprehensive analysis of three-dimensional genomic structures in human cells. These ranged from gastric cancer cell lines, patient samples, normal gastric epithelial cells, and EBV-associated gastric cancer. Combined with virus infection analyses, the researchers found abnormally activated genomic regions specific to EBV-positive stomach cancer. Experimental EBV infection of cultured stomach cells reproduced the phenomena of EBV binding to these inactive and closed genomic regions and their abnormal activation.
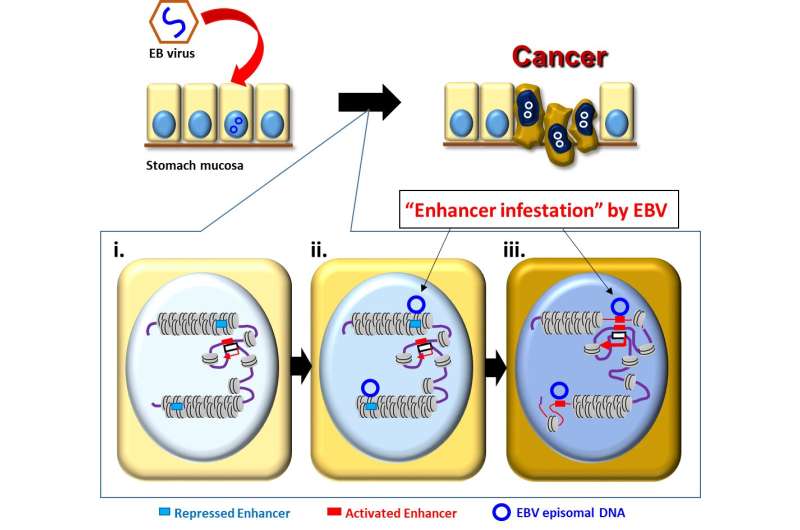
"Cells put active marks on genomic regions necessary for their behaviors and utilize them, and inactive marks on unnecessary genomic regions that are tightly closed and not to be utilized," explained Prof Kaneda. "We made the striking observation that strong inactive marks were lost in specific genomic regions when we infected stomach cells with EBV."
The researchers further found that genetic enhancers (short pieces of DNA that help encourage genes to make proteins) "silenced" in the closed regions were activated by the virus to upregulate nearby cancer-related genes, leading to the proliferation of cancerous cells. This "enhancer infestation" model, as the researchers termed it, reveals a novel mechanism of tumorigenesis that does not require genetic alterations, and instead works by reprogramming the epigenetic landscape of human cells to convert latent enhancers from a silenced to an active state.
Prof Patrick Tan, who is also a member of the Singapore Gastric Cancer Consortium, remarked, "In all EBV-positive stomach cancer cells and primary stomach cancer patient samples studied, EBV DNA bound to largely the same genomic regions that also showed abnormal activation. These same regions also changed from inactive to active states by experimental EBV infection."
This mechanism of "enhancer infestation" led to the activation of neighboring proto-oncogenes in human cells and it is likely to contribute to EBV-associated oncogenesis in multiple cancer cell types. Notably, the researchers also found that, even after eliminating EBV genomes, epigenetic modifications that were induced continued to persist, suggesting a 'hit-and-run' mechanism in which, once an EBV episome alters the chromatin topology of human cells, these altered topologies are stable and persist even after removal of the EBV episome.
Prof Kaneda reiterated, "While 8 to 10% of stomach cancer is associated with EBV, we believe our enhancer infestation model provides a new mechanism of cancer involving epigenomic alterations and viral infection that may be relevant to a broader range of cancers and associated diseases."
Prof Tan added, "Infections by EBV are estimated to cause over 200,000 cancers per year worldwide, including certain stomach cancers. Our study highlights new potential drug targets in EBV-positive malignancies, revealed by epigenetics and previously invisible using more conventional genetic sequencing studies."
More information: Okabe, A., Huang, K.K., Matsusaka, K. et al. Cross species chromatin interactions drive transcriptional rewiring in Epstein Barr virus positive gastric adenocarcinoma. Nature Genetics (2020). doi.org/10.1038/s41588-020-0665-7