Studying the human genome to understand the risk of autism

A team of researchers at CHU Sainte-Justine and Université de Montréal has succeeded in using bioinformatics to develop a statistical model to assess how the gain or loss of genetic material impacts the risk of autism.
The results of this work are presented today in the American Journal of Psychiatry.
This project highlights the fact that, besides the genetic mutations formally associated with autism, there are thousands of genes in the human genome that contribute to increasing the risk and severity of autism based on genetic scores.
A condition with many faces
Autism is a neurodevelopmental condition with many faces. It is defined by significant gaps in two areas, namely social communication and interaction on the one hand, and limited or repetitive behavior, activities and interests on the other hand.
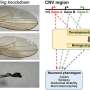
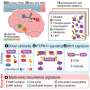
Deletions (loss) or duplications (gain) of a portion of the genome, also known as copy number variants (CNVs), are identified and suspected to cause the syndrome in up to 20% of individuals with autism.
"However, only a handful of CNVs could be characterized and attributed to autism. The majority of rare CNVs remain unknown and there's little information on their effect-size on cognitive and behavioral traits associated with autism," explained Dr. Sébastien Jacquemont, a genetics clinician-scientist at CHU Sainte-Justine and principal author of the study.
Assessing the effect of loss of genetic material
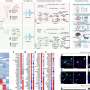
"We recently developed a statistical model to estimate the effect-size of CNVs on IQ based on genetic scores. We then sought to extend the model to autism susceptibility and to various cognitive measures such as social skills," explained Guillaume Huguet, Ph.D.
"The 'level of intolerance to gene mutation,' or if you prefer 'how mutation increases the risk of developing a disease,' best explains the effect of CNVs on IQ and the risk of autism," added Elise Douard, MSc, co-lead author of the study.
"Thanks to the statistical model, we were able to determine that effects of CNVs on IQ are twice as important for deletions than they are for duplications," she said. "Once CNV effects on IQ are accounted for, autism susceptibility remains mostly unchanged for duplications but decreases for deletions. This suggests that deletions are more specifically associated with intellectual disability than the risk of autism."
She added: "This model allows us to predict the risk of autism associated with any genome deletion or duplication based on genetic scores."
Findings are critical
"These findings are critical because unknown CNVs are diagnosed in a large proportion of patients," said Jacquemont. "Our online prediction tool will help health professionals rapidly evaluate the effect-size of CNVs on the risk of autism as well as on various cognitive, language and motor skills measures."
He added: "Knowing the nature of genetic variations in a given disease will allow us to better diagnose and treat patients to positively impact their life trajectory."
Neurodevelopmental disorders, including autistic disorders, are a major health issue throughout the world. In Canada, one in 68 children is diagnosed with an autistic disorder. This figure rises to one in 64 children in Quebec. These disorders are also associated with comorbidities: in 19 to 48 percent of cases, for example, autism is diagnosed in combination with an intellectual disability.
More information: Elise Douard et al. Effect Sizes of Deletions and Duplications on Autism Risk Across the Genome, American Journal of Psychiatry (2020). DOI: 10.1176/appi.ajp.2020.19080834