How transcription factors work together in cancer formation
A new study co-authored by University of Colorado Cancer Center researcher Srinivas Ramachandran, Ph.D., shows how DNA segments known as enhancers function in cells.
The paper published last month in Molecular Cell highlighted the work from Ramachandran, along with Satyanarayan Rao, both part of the Department of Biochemistry and Molecular Genetics at the CU School of Medicine, and Kami Ahmad from the Fred Hutchinson Cancer Research Center.
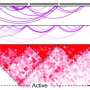
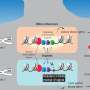
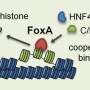
Enhancers are DNA sequences that drive cell-type-specific gene expression, developmental transitions, and cellular responses to external stimuli. They typically have multiple binding sites for transcription factors, which are proteins that help turn specific genes "on" or "off" by binding to nearby DNA. Ramachandran wanted to find out what the role of those multiple binding sites was in driving enhancer function, and if the transcription factors were binding to the multiple enhancer sites randomly or in a coordinated fashion (which in biology is called "cooperativity").
"Think of the two binding sites in the enhancer as two chairs at a table in a coffee shop. The chairs could be occupied by strangers. They arrive and leave the table at random times, and any time they overlap would be purely by chance. If two friends show up to the coffee shop, they will spend much more time together at the table compared to random strangers," he says.
"Biologically, the implication for transcription factors occupying enhancers at the same time is very big. Our starting assumption was that the transcription factors were more like strangers than friends at a coffee shop. We wanted to develop methods to first measure if there is any cooperativity, and if we found any, then ask how the cell might make the factors work together."
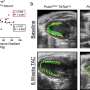
The researchers observed the binding process in the cells of fruit flies, developing two methods to see if multiple transcription cells were binding together or separately at two adjacent binding sites in enhancers.
"Then we could actually say, what is the expectation of them spending time together?" Ramachandran says. "If we know one is bound 50% of the time and the other is bound 50% of the time, then my expectation for no cooperativity is, by multiplying these probabilities, that two of them are together 25% of the time just by chance. But if two of them are together 40% of the time, that means there's some cooperativity. That somehow the enhancer is having conditions that make the transcription factors bind at the same time."
The researchers found that up to 60% of the enhancers they observed had at least one coordinated binding event, meaning the transcription factors were turning on and off together and working together to turn genes off and on. If scientists can better understand why that process happens, he says, they may be able to better understand how enhancers are coopted by diseases like cancer.
"It gets back to how enhancers evolved and how enhancers function," Ramachandran says. "If you think about cancer, it's cells changing their identity from normal cells. They are taking advantage of systems that exist but are usually silenced in that cell type. Understanding how enhancers work actually lets us understand how they are taken advantage of as cells change their identity during disease."
More information: Satyanarayan Rao et al. Cooperative binding between distant transcription factors is a hallmark of active enhancers, Molecular Cell (2021). DOI: 10.1016/j.molcel.2021.02.014