Model can predict how drug interactions influence antibiotic resistance
Scientists have proposed a modeling framework which could predict how antibiotic resistance will evolve in response to different treatment combinations, according to a study published in eLife.
The research, co-led by University of Michigan biophysicist Kevin Wood, could help doctors optimize the choice, timing, dose and sequence of antibiotics used to treat common infections, helping to halt the growing threat of antibiotic resistance to modern medicine.
"Drug combinations are a particularly promising approach for slowing resistance, but the evolutionary impacts of combination therapy remain difficult to predict, especially in a clinical setting," said first author Erida Gjini, a researcher at the University of Lisbon, Portugal.
"Interactions between antibiotics can accelerate, reduce or even reverse the evolution of resistance, and resistance to one drug might also influence resistance to another. These interactions involve genes, competing evolutionary pathways and external stressors, making it a complex scenario to pick apart."
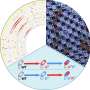
In their study, Gjini and Wood sought to simplify things. They took a fundamental measurement of microbe fitness—their growth rate, measured by a simple growth curve over time—and linked this to resistance to two theoretical drugs. In the model, they assumed that drug-resistant mutants respond to a high concentration of drug in exactly the same way that drug-sensitive cells respond to a low concentration of drug.
This rescaling assumption means that the growth behavior of mutants can be inferred from the behavior of the ancestral (sensitive) cells, simply by measuring their growth over a range of concentrations. The team then connected this assumption to a famous statistical relationship, called the Price equation, to explain how drug interactions and cross-resistance impact the way populations evolve resistance quantitatively and adapt to drug combinations.
This rescaling model showed that the selection of resistance traits is determined by both the drug interaction and by cross-resistance (where cells develop resistance to one of the drugs and become resistant to the second drug at the same time). A mixture of two drugs in the model leads to markedly different growth trajectories and rates of growth adaptation, depending on how the drugs interact.
For example, growth adaptation can be slowed by drugs that mutually weaken one another—drugs that interact "antagonistically"—but the effect can be tempered or even reversed if resistance to one drug is highly correlated with resistance to the other. The predictions of the model help explain counterintuitive behavior observed in past experiments, such as the slowed evolution seen when combinations of tigecycline and ciprofloxacin—two antibiotics commonly used in clinical settings—are applied simultaneously to the opportunistic pathogen E. faecalis.
Having established the basic model, the team then added in the effect of mutations on drug resistance. They looked at two different routes to accumulating mutations. In the first, there was a uniform pathway between the ancestral genetics and all possible mutation combinations. In the second, they assumed that mutations must arise in a specific sequence. They used a theoretical combination of two drugs, one at a higher dose than the other, and found that the sequential pathway leads to slower adaptation of growth, reflecting its evolution to the first fittest mutant before adapting further.
In addition to being able to include mutations in the model, they also tested whether they could predict the effects of different timings and sequences of antibiotic treatment. They studied two sequential regimes, A and B, based on different dosage combinations of tigecycline and ciprofloxacin. They found that both the resistance levels to the two drugs and the growth rate increases during treatment, as they anticipated. But the dynamics of this increase depends on the relative duration of each treatment and the total treatment length.
"We have built a model that incorporates drug interactions and cross-resistance to predict how microbes will adapt over time in a way that can then be experimentally measured," said Wood, U-M associate professor of biophysics and physics.
"In contrast to the classical genetics-based approaches to studying drug resistance, we used simple scaling assumptions—something commonly used in physics—to dramatically reduce the complexity of the problem. The approach helps us unravel a number of competing evolutionary effects and may eventually offer a framework for optimizing time-dependent, multidrug treatments."
More information: Ziah Dean et al, Antibiotic interactions shape short-term evolution of resistance in Enterococcus faecalis, Dryad (2020). DOI: 10.5061/dryad.j3tx95x92