'Digital twins,' an aid to give individual patients the right treatment at the right time
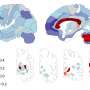
An international team of researchers have developed advanced computer models, or "digital twins," of diseases, with the goal of improving diagnosis and treatment. They used one such model to identify the most important disease protein in hay fever. The study, which has just been published in the open access journal Genome Medicine, underlines the complexity of disease and the necessity of using the right treatment at the right time.
Why is a drug effective against a certain illness in some individuals, but not in others? With common diseases, medication is ineffective in 40–70 percent of the patients. One reason for this is that diseases are seldom caused by a single "fault" that can be easily treated. Instead, in most diseases the symptoms are the result of altered interactions between thousands of genes in many different cell types. The timing is also important. Disease processes often evolve over long periods. We are often not aware of disease development until symptoms appear, and diagnosis and treatment are thus often delayed, which may contribute to insufficient medical efficacy.
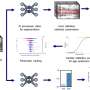
In a recent study, an international research team aimed to bridge the gap between this complexity and modern health care by constructing computational disease models of the altered gene interactions across many cell types at different time points. The researchers' long-term goal is to develop such computational models into "digital twins" of individual patients' diseases. Such medical digital twins might be used to tailor medication so that each patient could be treated with the right drug at the right time. Ideally, each twin could be matched with and treated with thousands of drugs in the computer, before actual treatment on the patient begins.
The researchers started by developing methods to construct digital twins of patients with hay fever. They used a technique, single-cell RNA sequencing, to determine all gene activity in each of thousands of individual immune cells—more specifically white blood cells. Since these interactions between genes and cell types may differ between different time points in the same patient, the researchers measured gene activity at different time points before and after stimulating white blood cells with pollen.
In order to construct computer models of all the data, the researchers used network analyses. Networks can be used to describe and analyze complex systems. For example, a football team could be analyzed as a network based on the passes between the players. The player that passes most to other players during the whole match may be most important in that network. Similar principles were applied to construct the computer models, or "twins," as well as to identify the most important disease protein.
In the current study, the researchers found that multiple proteins and signaling cascades were important in seasonal allergies, and that these varied greatly across cell types and at different stages of the disease.
"We can see that these are extremely complicated changes that occur in different phases of a disease. The variation between different times points means that you have to treat the patient with the right medicine at the right time," says Dr. Mikael Benson, professor at Linköping University, who led the study.
Finally, the researchers identified the most important protein in the twin model of hay fever. They show that inhibiting this protein, called PDGF-BB, in experiments with cells was more effective than using a known allergy drug directed against another protein, called IL-4.
The study also demonstrated that the methods could potentially be applied to give the right treatment at the right time in other immunological diseases, like rheumatism or inflammatory bowel diseases. Clinical implementation will require international collaborations between universities, hospitals and companies.
More information: Xinxiu Li et al, A dynamic single cell-based framework for digital twins to prioritize disease genes and drug targets, Genome Medicine (2022). DOI: 10.1186/s13073-022-01048-4