Genetic variation explains racial disparity in esophageal cancer cases
Esophageal adenocarcinoma (EAC) is a type of cancer affecting the mucus-secreting glands of the lower esophagus—the tube connecting the throat to the stomach. It is the most common form of esophageal cancer and often preceded by Barrett's metaplasia (BE), a deleterious change in cells lining the esophagus.
Though the cause of EAC remains unclear, cell mutations have been linked, perhaps induced by risk factors like tobacco or alcohol use or chronic damage caused by gastroesophogeal reflux disease or GERD. But the driver of these mutations has proven confounding, in part because the incidence of EAC is disproportionate: African Americans are approximately four- to five-fold less likely to develop EAC than Caucasians. They are also less likely to experience BE.
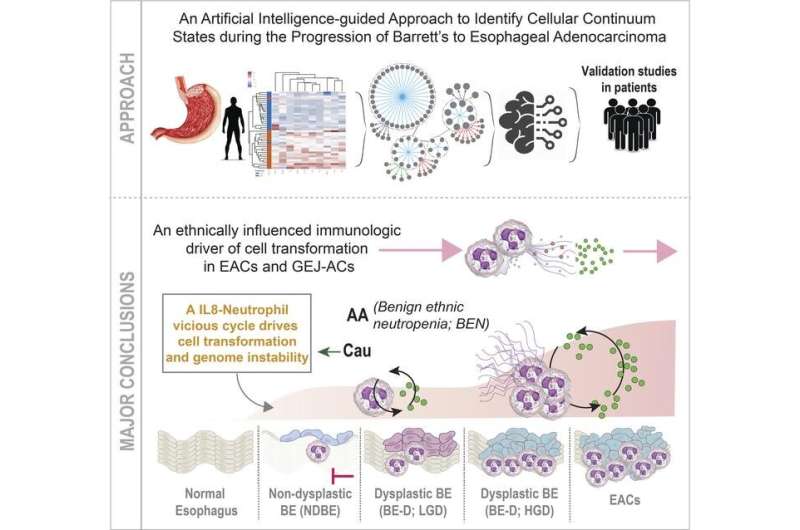
In a new study, published September 22, 2022 in the journal JCI Insight, researchers at the University of California San Diego School of Medicine, with colleagues in Brazil, used artificial intelligence-guided tools to pinpoint both a specific type of immune cell as the disease driver, and a specific genetic variation known as a SNP (single nucleotide polymorphism) that acts as a protective factor in African Americans.
SNPs represent a difference in a single DNA building block, called a nucleotide. They occur normally throughout a person's DNA. Most have no effect on health or development, but some are associated with disease when the variations are shared by many individuals who also share a predisposition to that disease.
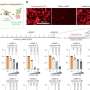
The team, led by co-corresponding authors Pradipta Ghosh, MD, professor in the departments of Medicine and Cellular and Molecular Medicine at UC San Diego School of Medicine, and Debashis Sahoo, Ph.D., associate professor in the departments of Pediatrics at UC San Diego School of Medicine and Computer Science at UC San Diego Jacobs School of Engineering, used artificial intelligence and machine learning to identify the progression from BE to EAC in different cell types and tissues, confirming their findings using organoids, patient-derived biopsies and a cross-sectional study of 113 persons with BE and EAC.
The work confirmed that all EACs originate from BE and pinpointed the role of a version of neutrophil, a white blood cell that acts as the immune system's first line of defense, as the driver of cellular transformation in both EACs and gastroesophageal junction adenocarcinoma, a rare esophageal cancer that occurs at the connection between the esophagus and stomach.
Both cancers carry poor prognoses, with an overall 5-year survival of less than 20 percent.
"This neutrophil driver was prominent in Caucasians, but notably absent in African Americans," said Sahoo. "Conversely, SNPs associated with ethnic changes in absolute neutrophil count, such as benign ethnic neutropenia characterized by lower numbers of neutrophils but no increased risk of infection, are common in persons of African ancestry and may act as a deterrent to prevent BE from becoming EAC."
The authors said the findings are important because they trace the cellular continuum from a precancer state (BE) to cancer, and clarify the roles of neutrophils and genetic variation by ethnicity.
"A central challenge in genetics is to understand how changes in DNA result in observable changes in an organism," said Ghosh. "In this instance, we found that a SNP that reduces the total number of circulating neutrophils in African Americans also protects them from EACs, a cancer whose progression is driven by neutrophils."
Ghosh and colleagues are cautiously optimistic that neutrophil targeted therapies may emerge as potential immunotherapies in EACs. She said researchers will continue to investigate these possibilities.
The study was executed by an international team of gastroenterologists, bioinformaticians, experts in pre-cancer biology and cancer genetics, assembled under the umbrella of the Institute for Network Medicine at UC San Diego School of Medicine. The institute fosters several transdisciplinary programs that use biological networks created with AI tools from the Center for Precision Computational Systems Network to chart unknown territories of disease.
More information: Pradipta Ghosh et al, AI-assisted discovery of an ethnicity-influenced driver of cell transformation in esophageal and gastroesophageal junction adenocarcinomas, JCI Insight (2022). DOI: 10.1172/jci.insight.161334