Genomic transposable elements modify the progression of Parkinson's disease
A study involving researchers from the University of Liverpool describes how transposable elements are associated with Parkinson's subtypes and impact disease trajectory.
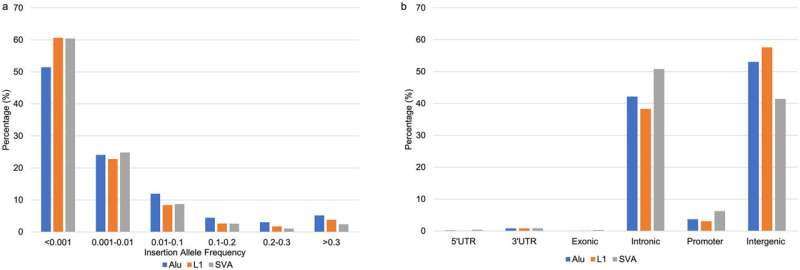
The study, published in Experimental Biology and Medicine, analyzed the variation of transposable elements—DNA sequences that have the ability to change their position within a genome—and their impact on different trajectories of Parkinson's disease.
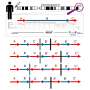
Researchers used patient data from the Parkinson's Progression Markers Initiative, one of the largest international longitudinal cohorts of Parkinson's patients, from patients who also had whole genome sequencing data available. This allowed researchers to develop a comprehensive genomic screen for the transposable elements in these patients and to detect their impact on disease progression.
The study found that transposable elements were associated with Parkinson's subtypes and disease trajectory. Some transposable elements in a patient's genome predicted faster progression of the disease, with very fast deterioration of motor or cognitive functions. Interestingly, the effect of transposable elements wasn't unidirectional. Depending on their type and position in the genome, these elements may have a negative or positive effect.
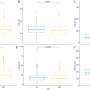
Some elements were associated with a strong and clinically meaningful change in functional Parkinson's scores, including a 40-point change in the Unified Parkinson's Disease Rating Scale. At the same time, some transposable elements were protective and associated with a slower disease progression suggesting a slowing of neuronal loss and neurodegeneration.
Transposable elements make up more than 70% of the human genome, but until recently, they have been considered as "junk" DNA without any meaningful function. These elements have been largely ignored in most genetic studies and could therefore contribute to the currently unknown genetic factors that influence disease development.
Previous studies from these researchers have shown that these elements have a significant regulatory role in the genome. They regulate the expression of many genes, orchestrate complex genetic activation where multiple genes are needed for a complex response, and the growing body of evidence suggests that these elements have a major pathogenetic effect, with several elements shown to cause disease.
The study was led by Professor Sulev Koks, head of Genetic Epidemiology Research at Perron Institute for Neurological and Translational Science and the Center for Molecular Medicine and Innovative Therapeutics at Murdoch University in Perth, Western Australia, with colleagues at Perron Institute, Murdoch University and Professor John Quinn's group at the University of Liverpool.
"Transposable elements are part of the genome which is known as the 'dark genome'", senior author Professor Sulev Koks said. "Our study showed that the dark genome may have a much more significant impact on the pathophysiology of complex disease than previously estimated."
"The elements inside the dark genome could enhance or slow the progression of Parkinson's disease and therefore open up new opportunities for precision medicine."
The University of Liverpool's Professor John Quinn said, '"Our body of work over a decade highlights the importance of targeting these elements for therapeutic intervention in the progression of neurodegenerative diseases which is the subject of our current work not only in Parkinson's Disease but also Motor Neuron Disease. "This could include repositioning of drugs from other disease areas in which these elements are implicated in disease progression."
More information: Sulev Kõks et al, Non-reference genome transposable elements (TEs) have a significant impact on the progression of the Parkinson's disease, Experimental Biology and Medicine (2022). DOI: 10.1177/15353702221117147