This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
CRISPR studies identify many promising therapeutic targets for multiple myeloma
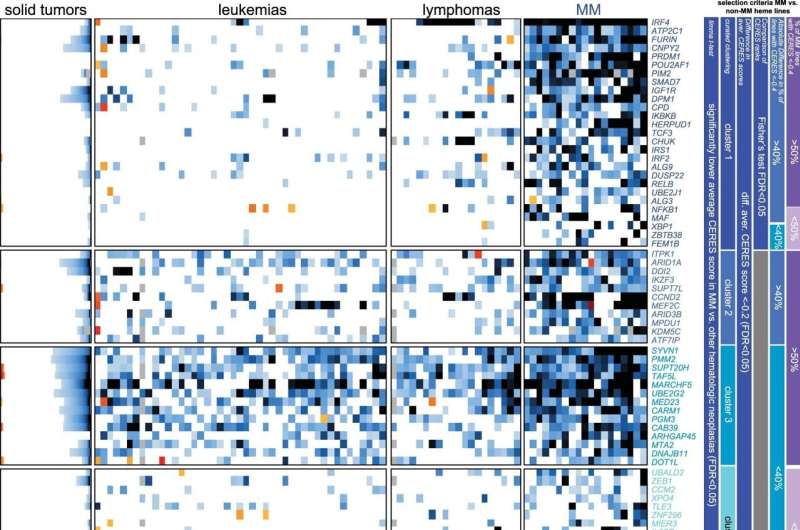
New research from Dana-Farber Cancer Institute has identified 116 genes as key molecular vulnerabilities for multiple myeloma. Most of these genes are potential leads for the discovery of new therapies for this disease. The study was published in Nature Cancer.
Multiple myeloma is an incurable blood cancer that affects plasma cells, the antibody-producing cells of the body. To identify the leads, the team studied cell lines from multiple myeloma and a large collection of non-multiple myeloma tumor types. Using CRISPR to disable genes one-by-one in each cell line, they identified genes that are essential for the survival of the multiple myeloma cells but much less important for cancer cells from other lineages.
While the findings point to a range of potential targets for myeloma therapies, they also suggest a different approach to targeted therapy in general—one in which therapies take aim at those unique functional vulnerabilities that distinguish an individual type of cancer from all others, regardless of whether these vulnerabilities are caused by genetic mutations.
"It's very exciting to find so many new and underappreciated candidate targets," says Dana-Farber researcher Constantine Mitsiades, MD, Ph.D., who led the study. "We are actively working to determine which of these can be targeted by new therapeutic interventions for multiple myeloma."
Defining functional vulnerabilities that distinguish one cancer type from others
For nearly two decades, the predominant approach to finding candidate drug targets has started with a search for mutated or overexpressed genes associated with a type of cancer compared with its respective normal cells. These techniques have led to targeted cancer therapies that have benefited many patients.
In this study, however, the researchers sought to identify the distinct functional vulnerabilities that distinguish myeloma from other cancers—no mutations or alterations needed.
"This study emphasizes how selectively critical a gene is for myeloma cells," says Mitsiades. "If turning off a gene causes important damage in multiple myeloma cells, but not all or most other cell types, we've found something important."
The team relied on multiple analytical tools to narrow down a list of these important genes that are preferentially essential for multiple myeloma cells. The list includes genes that make molecules that control which genes are activated or how proteins are secreted in plasma cells, and many other genes not previously considered important for the biology of myeloma or most other cancers. The team also validated the large majority of these genes were essential for multiple myeloma cells in animal models.
This approach to discovery of new therapeutic targets, done in collaboration with the Broad Institute of MIT and Harvard, identified an almost completely new and unique set of potential multiple myeloma cell vulnerabilities compared to previous studies. For example, of the 116 genes identified, only a few had been found in earlier studies looking for mutations or alterations that contribute to multiple myeloma.
"It's a shift in thinking about targeted therapies for a given cancer," says Mitsiades, "Instead of targeting the genes that are mutated more frequently or expressed at higher levels, we're thinking about molecular targets that are more functionally important for this specific cancer type."
Past and future of targeting functional vulnerabilities for multiple myeloma
A few of the genes identified in this study are already known drug targets for multiple myeloma. The multiple myeloma drug thalidomide (and drugs derived from it), for example, degrades proteins made by the genes IKZF3 and IKZF1, a discovery made ten years ago by co-author Benjamin L. Ebert, MD, Ph.D., chair of Medical Oncology at Dana-Farber, and Nobel laureate William G. Kaelin, Jr., MD, Sidney Farber Professor of Medicine at Harvard Medical School and Dana-Farber. Those two genes showed up on Mitsiades' list.
"It was reassuring that molecules that are already known as important therapeutic targets for multiple myeloma and not for most other cancers came up as preferential targets for myeloma in our studies," says Mitsiades. "It strongly supports the value of the other leads we identified."
Therapeutics that target molecules that are preferentially essential for multiple myeloma but not for most other cancer types might have another advantage. They could also be less likely to harm the normal tissues that correspond to these other cancer types. In turn, this could mean lower probability for serious side effects. Further, they could work for many people with multiple myeloma because their tumor cells do not need have to have a certain mutation in order to respond to such treatments.
More information: Ricardo de Matos Simoes et al, Genome-scale functional genomics identify genes preferentially essential for multiple myeloma cells compared to other neoplasias, Nature Cancer (2023). DOI: 10.1038/s43018-023-00550-x