This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Cancer researchers discover optimal cancer-killing T cells
A team of cancer researchers, led by the University of Houston, has discovered a new subset of T cells that may improve the outcome for patients treated with T-cell therapies.
T cell-based immunotherapy has tremendous value to fight, and often eliminate, cancer. The strategy activates a patient's immune system and engineers a patient's own T cells to recognize, attack and kill cancer cells. In this way, the body's own T cells become living drugs.
While T-cell immunotherapy has revolutionized cancer treatment, there is still much to learn. Unfortunately, not all patients respond to these therapies, so a better understanding of the properties of engineered T cells is necessary to improve clinical responses.
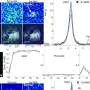
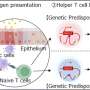
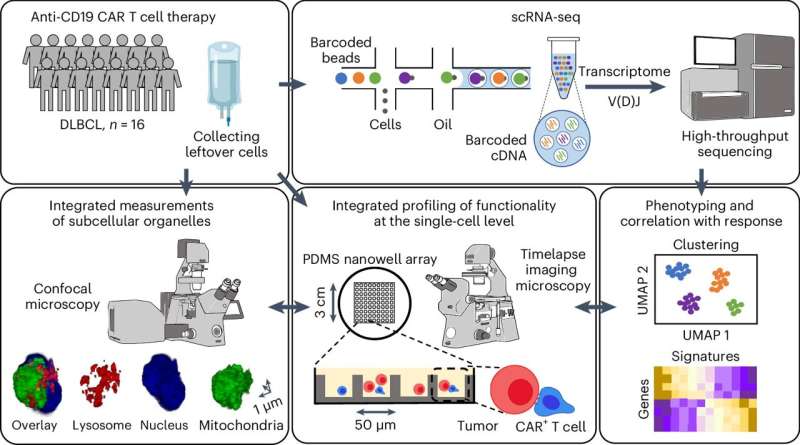
One such study is reported in Nature Cancer by the laboratory of Navin Varadarajan, M.D. Anderson Professor in the William A. Brookshire Department of Chemical and Biomolecular Engineering. The study uses the patented TIMING (Timelapse Imaging Microscopy in Nanowell Grids) approach which applies visual AI to evaluate cell behavior, movement and ability to kill.
"Our results showed that a subset of T cells, labeled as CD8-fit T cells, are capable of high motility and serial killing, found uniquely in patients with clinical response," reports first author and recent UH graduate Ali Rezvan.
To discover the CD8-fit cells, the team used TIMING to track interactions between individual T cells and tumor cells across thousands of cells and integrated the results with single-cell RNA sequencing data.
"Chimeric antigen receptors (CAR) T cells used for the treatment of B cell malignancies can identify T-cell subsets with superior clinical activity. Using infusion products of patients with large B cell lymphoma, we integrated functional profiling using TIMING with subcellular profiling and scRNA-seq to identify a signature of multifunctional CD8 T cells (CD8-fit)," said Rezvan.
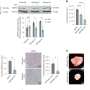
"We profiled these cells using single-cell RNA sequencing to identify the CD8-fit molecular signature that could be used to predict durable patient outcomes to T-cell therapies and validated our findings with independent datasets."
The team also found that the CD8-fit signature is present in pre-manufactured T cells, longitudinally persists in patients post-infusion, and most importantly, is associated with long-term positive clinical responses. According to the researchers, it is likely that these T cells can drive clinical benefit in other tumors.
"This work illustrates the excellence of graduate students Ali Rezvan and Melisa Montalvo; and post-doctoral researchers Melisa Martinez-Paniagua and Irfan Bandey among others," said Varadarajan.
In addition to the UH team, collaborators include Sattva Neelapu and Harjeet Singh, The University of Texas MD Anderson Cancer Center, Houston; Mike Mattie, Kite Pharma; Nabil Ahmed, Texas Children's Hospital, Baylor College of Medicine, Houston; and Mohsen Fathi, CellChorus.
More information: Ali Rezvan et al, Identification of a clinically efficacious CAR T cell subset in diffuse large B cell lymphoma by dynamic multidimensional single-cell profiling, Nature Cancer (2024). DOI: 10.1038/s43018-024-00768-3