This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Unlocking key answers on cell functioning for improved cancer treatments
Peter Mac researchers have found the answer to a decades-long question on cell functioning that could lead to improved cancer treatments in the future.
Every cell in the human body has the same DNA, but different cells do different jobs. This research, published in Nature Genetics, is helping to explain how this is possible and the implications could be vast. Professor Mark Dawson, a physician-scientist and Associate Director of Research at Peter Mac, said he is excited that we now have new insights that better explain how a cell's fate is determined.
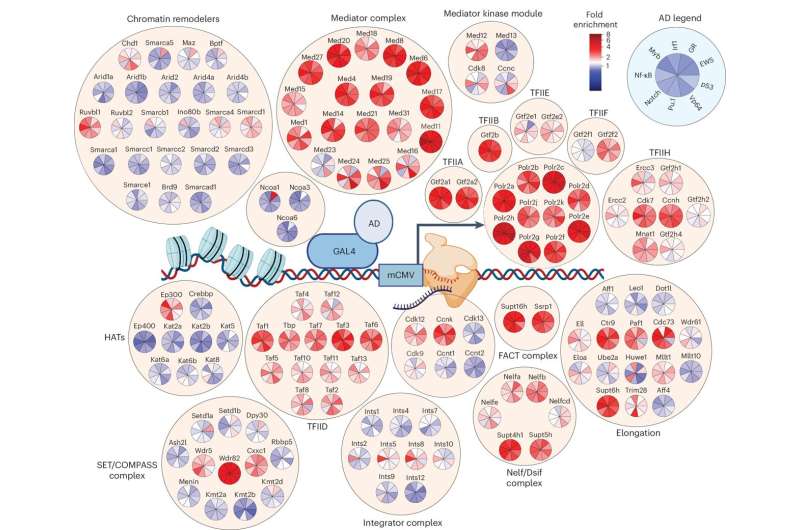
"The way a cell functions is the result of 'transcription factors' that scan our DNA and determine which genes should be turned on and by how much," he said.
"We looked at how these transcription factors recruit and deliver the machinery required for genes to be turned on. Until now, we did not know how the 'transcription factors' choose the right machinery for a gene to be read and expressed.
"This has been a decades-long question, and we are excited to have helped solve part of the problem as this knowledge on exactly how transcription factors make decisions around which machinery is required to activate a gene provides us with fundamental knowledge on life."
The study uncovered that transcription factors choose a unique compendium of components to drive gene expression to create the desired action whether it be controlling a cell's energy consumption, launching an immune response or some other function our body requires. Professor Dawson said that we can liken this to how cars are built and explained how this important discovery is key to finding better treatments for a range of diseases.
"A F1 racing car is very different to the family people mover or even a tractor, some cars are designed to move very fast, others to transport precious cargo and some are the work horse," he said.
"We uncovered that the same is true for gene expression and this is determined by the components recruited by transcription factors. They can determine which genes can rapidly change, for example, when we need to fight infection and we need a quick response or which genes need to be slow and steady pumping out messages required for cellular housekeeping function.
"This insight into how transcription factors can nuance gene expression is incredibly important, and we hope to harness this to help us treat a range of diseases in the future.
"If we think about cancer, the mutations in a cancer can stop the transcription factor from choosing the right components for the gene to be expressed correctly, it is like jumbling up parts of the car so it no longer drives reliably."
Dr. Charles Bell, post-doctoral researcher at Peter Mac, said they developed a platform to screen the function of thousands of components used by transcription factors to determine how a gene is expressed.
"We will now use this platform to understand other processes linked to the gene expression," he said.
"The answers to these questions will help us find new ways to treat not just cancer but many other diseases in the future."
More information: Charles C. Bell et al, Comparative cofactor screens show the influence of transactivation domains and core promoters on the mechanisms of transcription, Nature Genetics (2024). DOI: 10.1038/s41588-024-01749-z