Immune system development linked to leukemia
Scientists have discovered a genetic signature that implicates a key mechanism in the immune system as a driving force for a type of childhood leukaemia.
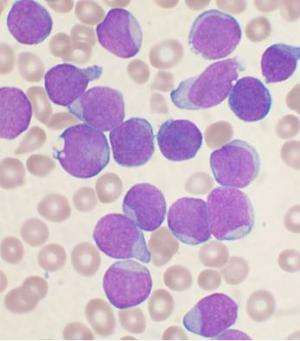
Acute lymphoblastic leukaemia or ALL is the most common form of childhood leukaemia. A key factor driving this leukaemia for one in four ALL patients is a mutation that causes two of their genes, ETV6 and RUNX1, to fuse together. This genomic alteration happens before birth and kick starts the disease. But on its own the fusion gene cannot cause cancer; it requires additional mutations before the leukaemia can fully evolve and prompt symptoms.
The new study exploring how this process occurs was carried out by researchers from the Wellcome Trust Sanger Institute and The Institute of Cancer Research, London, with funding from the Kay Kendall Leukaemia Fund, Leukaemia and Lymphoma Research and the Wellcome Trust.
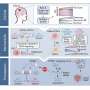
RAG proteins rearrange the genome in normal immune cells in order to generate antibody diversity. In ALL patients with the fusion gene, the team showed that these proteins can also rearrange the DNA of genes involved in cancer, leading to the development of leukaemia.
"For the first time, we see the combined events that are driving this treatable but highly devastating disease," says Dr Elli Papaemmanuil, first author from the Wellcome Trust Sanger Institute. "We now have a better understanding of the natural history of this disease and the critical events from the initial acquisition of the fusion ETV6-RUNX1 to the sequential acquisition of RAG-mediated genome alterations that ultimately result in this childhood leukaemia."
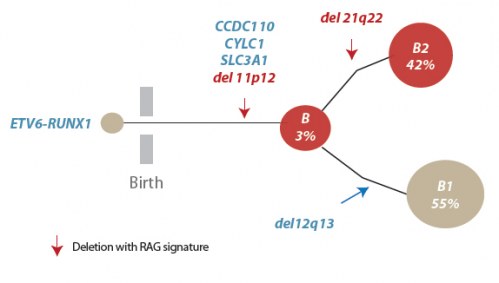
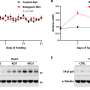
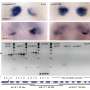
The team sequenced the genomes of 57 ALL patients with the fusion gene and found that genomic rearrangement and specifically deletions of DNA segments were the predominant drivers of the cancer. All samples showed evidence of events involving the RAG proteins.
The RAG proteins use a unique sequence of DNA letters as a signpost to direct them to antibody regions. The team found that remnants of this unique sequence lay close to more than 50 per cent of the cancer-driving genetic rearrangements. Importantly, this process often led to the loss of the very genes required for control of normal immune cell development.
It is the deletion of these genes that, in combination with the fusion gene, leads to the leukaemia. This striking genetic signature linking the RAG proteins to genomic instability is not found in other common cancers such as breast, pancreatic and prostate cancer, or other types of leukaemia.
"As we sequence more and more cancer genomes, we are increasingly understanding the mutational processes that underpin cancer's evolution," says Dr Peter Campbell, co-lead author from the Wellcome Trust Sanger Institute. "In this childhood leukaemia, we see that the very process required to make normal antibodies is co-opted by the leukaemia cells to knock-out other genes with unprecedented specificity."
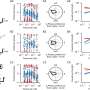
To better understand the genetic events that led up to the development of this cancer, the team used single-cell genomics, a state-of-the-art technique that looks at the DNA from an individual cell. Using samples from two patients, they were able to show that this cancer-causing process occurs many times and results in a continuous diversification of the leukaemia.
"It may seem surprising that evolution should have provided a mechanism for diversifying antibodies that can collaterally damage genes that then contribute to cancer," said Professor Mel Greaves, co-senior author of the study from The Institute of Cancer Research, London, "But this only happens because the fusion gene (ETV6-RUNX1) that initiates the disease 'traps' cells in a normally very transient window of cell development where the RAG enzymes are active, teasing out their imperfect specificity."
The team will now investigate how the RAG-mediated genomic instability accrues in cells with ETV6-RUNX1 fusion gene and what the role of this process is in the patients that relapse.
"The more we understand about the genetic events that underlie leukaemia and other cancers, the better equipped we are to develop improved diagnostics and targeted therapy for patients with this disease," adds Dr Campbell.
More information: Elli Papaemmanuil et al (2013). 'RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia' Advanced online publication in Nature Genetics 12 Jan. DOI: 10.1038/ng.2874