Bacteria-killing proteins cover blood type blind spot
A set of proteins found in our intestines can recognize and kill bacteria that have human blood type molecules on their surfaces, scientists at Emory University School of Medicine have discovered.
The results were published online Feb. 14 and are scheduled to appear in the journal Nature Medicine.
Many immune cells have receptors that respond to molecules on the surfaces of bacteria, but these proteins are different because they recognize structures found on our own cells, says senior author Richard D. Cummings, PhD, professor and chair of the Department of Biochemistry. "It's like having a platoon in an army whose sole purpose is to track down enemy soldiers that are wearing the home country's uniforms."
Blood type comes from differences in sugar molecules attached to proteins on red blood cells. If incompatible blood types are mixed, the antibodies from one person will make red blood cells from the other person clump together, with devastating results in an emergency. But someone's immune system usually doesn't make antibodies to the sugar molecules on his or her own red blood cells. That creates a potential blind spot that bacteria could exploit.
For example, a strain of E. coli (O86) has molecules on its surface like those in humans with blood type B. People with blood type B are unable to produce antibodies against E. coli O86. Although O86 is known to infect birds, it's not a major danger like other types of E. coli, some of which can cause severe diarrhea.
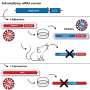
Cummings and his colleagues wanted to know why more bacteria haven't adopted the tactics of E. coli O86 to get around the immune system. Searching for proteins that could bind to the sugar molecules characteristic of blood types A and B, graduate students Sean Stowell, PhD, and Connie Arthur identified proteins called galectin-4 and galectin-8.
"These proteins are separate from antibodies and other parts of the immune system," Cummings says. "They kill bacteria like E. coli O86 all by themselves within a couple of minutes."
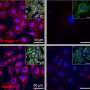
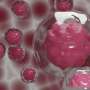
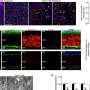
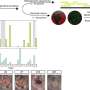
When E. coli O86 is exposed to these proteins and viewed by electron microscopy, "it looks as if somebody is tearing away at their outer membranes," he adds.
However, galectins-4 and -8 did not kill human red blood cells expressing blood group antigens. High levels of lactose (milk sugar) can inhibit the lethal activity of these galactins, whereas sucrose (cane sugar) does not.
"This raises the question of whether there are dietary effects, as from milk sugars or other dietary polysaccharides, that might inhibit activity of these galectins on intestinal microbes and their proliferation and colonization," Cummings says.
Cummings notes the unique properties of galectins-4 and -8 may provide an explanation for why the human population has such a diversity of sugar molecules on blood cells. The diversity may ensure that some part of the population might be able to fend off a bacterial infection. For example, ABO blood type seems to affect susceptibility to Helicobacter pylori, a bacterium linked to ulcers.
Galectins were thought to have evolved long before "adaptive immunity," the part of vertebrates' immune systems that is responsible for producing a variety of antibodies. Galectins may have allowed the generation of a diverse group of blood type sugar molecules in human tissues as a safe set of molecules to evolve because immunity is backstopped by galectins, Cummings says.
Galectins-4 and-8 were also able to kill another variety of E. coli that display a sugar molecule found on many mammalian cells, although more protein was needed. That leads to a question Cummings and his colleagues are investigating now: What else do galectins recognize, and how does that constrain the kinds of bacteria that can live in our intestines? In addition, it may now be possible, given these results, to engineer molecular changes in these galectins to allow them to kill other types of pathogenic bacteria that display other types of sugar molecules on their surface. Such developments could lead to new types of antibiotics for pathogenic microbes.
More information: S.R. Stowell et al. Innate immune lectins kill bacteria expressing blood group antigen. Nat. Med. 16, (2010).