Gene variations associated with risk of type 2 diabetes
For individuals of white European descent, certain variations of the gene HMGA1 are associated with type 2 diabetes mellitus, according to a study in the March 2 issue of JAMA.
Type 2 diabetes mellitus (DM) is a common metabolic disorder that affects nearly 250 million people worldwide, and is associated with major diabetes-related complications, including retinopathy, kidney disease and cardiovascular disease. Insulin resistance in muscle, liver, and fat tissue is a major feature of most patients with type 2 DM. There is considerable evidence that heredity is a major contributor to the insulin resistance of type 2 DM, according to background information in the article. "However, despite extensive investigations, including studies of candidate genes and the recent genome-wide association studies, the common genetic causes of insulin resistance remain elusive," the authors write. The researchers previously found that the protein HMGA1 is a key regulator of insulin receptor (INSR) gene expression.
Antonio Brunetti, M.D., Ph.D., of the University of Catanzaro, Catanzaro, Italy and colleagues conducted a study to examine the association of HMGA1 gene variants with type 2 DM, and included patients with type 2 DM and controls from 3 populations of white European ancestry. Italian patients with type 2 DM (n = 3,278) and 2 groups of controls (n = 3,328) were attending outpatient clinics and other health care sites in Calabria, Italy, during 2003-2009; U.S. patients with type 2 DM (n = 970) were recruited in Northern California clinics between 1994 and 2005 and controls (n = 958) were senior athletes without DM evaluated in 2004 and 2009; and French patients with type 2 DM (n = 354) and healthy controls (n = 50) were enrolled at the University of Reims in 1992. Genomic DNA was either directly sequenced or analyzed for specific HMGA1 mutations.
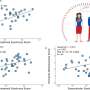
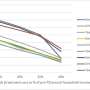
The researchers found that the most frequent functional HMGA1 variant, IVS5-13insC, was present in 7 percent to 8 percent of patients with type 2 DM in all 3 populations. The prevalence of this variant was higher among patients with type 2 DM (nearly 16 times higher odds of having this variant) than among controls in the Italian population (7.23 percent vs. 0.43 percent in one control group; and 7.23 percent vs. 3.32 percent in the other control group). In the U.S. population, the prevalence of IVS5-13insC variant was 7.7 percent among patients with type 2 DM vs. 4.7 percent among controls; in the French population, the prevalence of this variant was 7.6 percent among patients with type 2 DM and 0 percent among controls. In the Italian population, 3 other functional variants were observed. When all 4 variants were analyzed, HMGA1 defects were present in 9.8 percent of Italian patients with type 2 DM and 0.6 percent of controls.
"We believe our observation that nearly 10 percent of individuals with type 2 DM have deleterious variations in the gene encoding HMGAl has important clinical implications. First, the presence of these variants could serve as an early predictive marker of both insulin resistance and type 2 DM, especially in those individuals who have a family history of type 2 DM and related conditions. Second, the presence of these variants may predict the response to therapy. … Third, individuals who have functional HMGA1 variants and type 2 DM may have a different clinical course than other patients with type 2 DM, including differences in the development of macrovascular and microvascular complications. Fourth, the search for new therapies for type 2 DM could include agents that upregulate the expression of HMGA1," the authors write.
"In conclusion, our results indicate that variants in the HMGA1 gene are associated with type 2 DM in individuals of white, European descent. Further studies of the HMGAl gene and its variants, including studies in other racial types, are needed to understand the role of HMGA1 in insulin resistance and type 2 DM."
More information: JAMA. 2011;305[9]903-912.