Study pinpoints a genetic cause of most lethal brain tumor, may lead to new treatment
Researchers at Columbia University Medical Center (CUMC) have discovered that some cases of glioblastoma, the most common and aggressive form of primary brain cancer, are caused by the fusion of two adjacent genes. The study also found that drugs that target the protein produced by this genetic aberration can dramatically slow the growth of glioblastomas in mice. The findings were published today in the online edition of the journal Science.
"Our findings are doubly important," said study leader Antonio Iavarone, MD, professor of pathology and neurology at CUMC, and a member of the Herbert Irving Comprehensive Cancer Center (HICCC) at NewYork-Presbyterian Hospital/Columbia University Medical Center. "From a clinical perspective, we have identified a druggable target for a brain cancer with a particularly dismal outcome. From a basic research perspective, we have found the first example of a tumor-initiating mutation that directly affects how cells divide, causing chromosomal instability. This discovery has implications for the understanding of glioblastoma as well as others types of solid tumors."
The fusion of these two genes was observed in just three percent of tumors studied, so any therapy based on this particular genetic aberration would apply to only a small subset of glioblastoma patients. "It's unlikely that we will find a gene fusion responsible for most glioblastomas. But we may be able to discover a number of other gene fusions, each accounting for a small percentage of tumors, and each with its own specific therapy," said co-senior author Anna Lasorella, MD, associate professor of pathology and pediatrics at CUMC and a member of the Columbia Stem Cell Initiative and the HICCC.
"This is a very exciting advance in our understanding of cancer, and perhaps a first step toward a personalized, precision approach to the treatment of glioblastoma," said Stephen G. Emerson, MD, PhD, director of the HICCC and the Clyde '56 and Helen Wu Professorship in Immunology at the Columbia University College of Physicians and Surgeons.
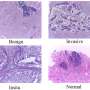
Glioblastomas are tumors that arise from astrocytes, star-shaped cells that make up the brain's supportive tissue. Since astrocytes reproduce quickly and are supported by a large network of blood vessels, glioblastomas are usually highly malignant. It is estimated that these tumors affect about 10,000 people in the United States each year. Glioblastoma is typically treated with surgery, followed by radiation and chemotherapy. However, the disease is invariably fatal, with a median survival of about 14 months after diagnosis, even with aggressive therapy. Glioblastomas took the lives of Senator Edward Kennedy in 2009 and New York Mets all-star catcher Gary Carter in 2012.
Several common single-gene alterations have been observed in glioblastoma. "However, therapies targeting these alterations have not improved clinical outcomes, most likely because they have systematically failed to eradicate the proteins to which the tumor is 'addicted,'" said Dr. Iavarone.
Dr. Iavarone and his colleagues suspected that glioblastomas might be addicted to proteins produced by gene fusions. Such fusions have been implicated in other cancers, notably chronic myelogenous leukemia (CML). Novartis AG's (NYSE: NVS) drug Gleevec (imatinib), which targets a fusion protein responsible for CML, has proved to be highly effective in arresting the disease.
In the current study, the CUMC researchers conducted genetic analyses of glioblastomas from nine patients, looking specifically for gene fusions. The most common fusion they observed involved the genes FGFR (fibroblast growth factor receptor) and TACC (transforming acidic coiled-coil).
Although each gene plays a specific role in the cell, sometimes errors in the DNA cause two ordinary genes to fuse into a single entity, with novel characteristics that can lead to a tumor," said co-senior author Raul Rabadan, PhD, assistant professor in the department of Biomedical Informatics and the Center for Computational Biology and Bioinformatics, Columbia Initiative in Systems Biology.
"We developed a new method for analyzing the cell's genomic material," he said. "First we looked at pieces of the glioblastoma genome from several samples, and then we extended the analysis to a large set of glioblastomas from the Cancer Genome Atlas project, sponsored by the National Cancer Institute."
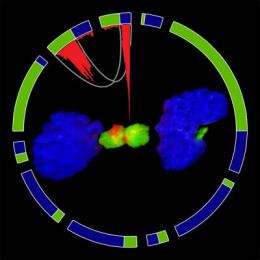
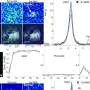
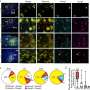
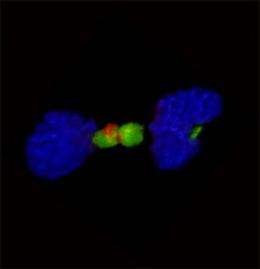
The researchers discovered that the protein produced by FGFR-TACC acts by disrupting the mitotic spindle, the cellular structure that guides mitosis (the division of a cell into two identical daughter cells). "If this process happens incorrectly, you get uneven distribution of the chromosomes. This condition, which is known as aneuploidy, is thought to be the hallmark of tumorigenesis," said Dr. Iavarone.
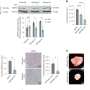
When FGFR-TACC was introduced into the brain cells of healthy mice, aggressive brain tumors developed in 90 percent of the animals, confirming that this gene fusion can lead to glioblastoma.
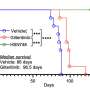
In another experiment, mice with this form of glioblastoma were given a drug that inhibits FGFR kinase, an enzyme essential for the protein produced by FGRF-TACC to do its work. The drug was found to prevent abnormal mitosis and double survival time, compared with a control group of mice that did not receive the drug.
Dr. Iavarone is currently establishing a cooperative study group, including CUMC and other brain tumor centers around the country, to conduct the trials of FGFR kinase inhibitors. Preliminary trials of these drugs (for treatment of other forms of cancer) have shown that they have a good safety profile, which should accelerate testing in patients with glioblastoma.
"This work is the result of an ongoing collaboration between a traditional and a computational lab. The synergy between the two approaches allows us to tackle complex biological problems in a high throughput fashion, providing a global view to the genome of glioblastoma," said Dr. Rabadan.
More information: "Transforming Fusions of FGFR and TACC Genes in Human Glioblastoma." Science, 2012.