Epigenetics: Neurons remember because they move genes in space
How do neurons store information about past events? In the Nencki Institute of Experimental Biology of the Polish Academy of Sciences in Warsaw, a mechanism unknown previously of memory traces formation has been discovered. It appears that at least some events are remembered thanks to... geometry.
Neurons are the most important cells of the nervous system. Scientists from the Nencki Institute of Experimental Biology of the Polish Academy of Sciences in Warsaw have shown that during neuron stimulation permanent changes are observed with respect to genes' arrangement within the cell nucleus. This discovery, reported in the Journal of Neuroscience, one of the most prestigious journals in the field of neurobiology, is significant for developing a better understanding of the processes going on in the mind and disorders of the nervous system, especially the brain.
"While conducting experiments on rats after epileptic seizures we have observed that a gene may permanently move deeper into the neuron's cell nucleus. Since modification of the geometrical structure of the nucleus leads to changes in gene expression, this is how the neuron remembers, what happened", explains Prof. Grzegorz Wilczyński from the Laboratory of Molecular and Systemic Neuromorphology at the Nencki Institute.
Neurons connect with each another via synapses, forming extended networks. In order for the neuronal networks to retain traces of stimuli which caused activation, the shape and functioning of individual synapses has to change. If stimulus trace is to be permanent, changes are necessary in the expression of many genes located in the cell nucleus of individual neurons.
Genes are sections of the deoxyribonucleic acid (DNA) chain coding proteins. But the presence of a gene in the DNA does not mean that it is active. It has been known for the past several years that gene expression also depends on the environment within the cell. Chromatin which fills cells contains gene activating or supressing substances.
"This somewhat resembles interpersonal relations. When you attend a social gathering the importance of what you say will have a different impact depending on the environment. If the environment is favourable, your opinion will be seized on and reinforced and you will achieve social impact. If the environment is less friendly, your opinion will be silenced", explains Prof. Wilczyński.
In the case of neurons the epigenetic processes during which gene expression is decided by the environment, to date have been associated only with chemical reactions within the chromatin. Research done at the Nencki Institute has shown that in neurons we deal with yet another type of epigenetic effects: changes to the spatial structure of the cell's nucleus resulting in the formation of permanent memory traces. This is possible for two reasons. First of all because of the presence of the nuclear membrane: genes can attach or detach from it, which impacts their expression. The second reason is related to the specific structure of the cell nucleus.
The nucleus of a cell consists of many globules, called chromosome domains or territories. Each domain is filled by just one chromosome, which may slightly move within its territory. As a result of such movement at the meeting points of the neighbouring domains, fragments of the DNA chains containing the different genes can come in contact. This leads to silencing of a group of genes or to their expression: formation of a transcription factory. However, a slight movement of the DNA chain in any domain changes the situation: the silenced gene unit resumes activity or the factory stops functioning.
Changes to the spatial arrangement of genes within the cell nucleus have already been observed in certain types of cells, among other in epithelium cells. Research done at the Nencki Institute has shown that external stimuli may cause changes within neurons. Moreover scientists proved that such changes are permanent and create a distinct genetic memory trace within the neuronal structure – despite no changes recorded in the DNA chains themselves.
Neurons used in this study came from rats after epileptic seizure, which is a brain plasticity disorder. During the seizure the activated neurons are places of turbulent gene expression. Scientists from the Nencki Institute decided to investigate two genes, known as BDNF and TRKB. In collaboration with the group of Prof. Marek Świtoński from the University of Life Sciences in Poznań, and with Prof. Marion Cremer from Munich, these genes' location within the DNA chains has been marked using a substance glowing after stimulation with laser light. Such preparations of neurons from control rats and neurons coming from rats after epileptic seizures were analysed under confocal microscope.
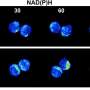
"Confocal microscope registers images only in the vicinity of its focal plane. Therefore each image represents a sort of flat, thin cross section through the preparation. To reconstruct from a set of many such slices the spatial structure of the cell nucleus and the arrangement of genes, we needed to design special software. This task turned out to be difficult since we were working at the limit of the microscope's resolving power", says Dr Błażej Ruszczycki from the Nencki Institute.
The software took one year to develop. It was used to study more than 5000 cell nuclei to determine the location of both genes of interest with relation to the centres of the nuclei and the nuclear membrane. For the BDNF gene, a change has been observed in its location of a few hundred nanometres (one billionth part of a metre); in the control animals this gene was present near the nuclear membrane or on it in 50% of the nuclei, while in animals after seizures this value dropped to approximately 25%.
"A double drop is a great change in biology. Moreover, we have observed that it remains visible for up to several weeks. The conclusion is therefore clear: past events are remembered by neurons also thanks to changes within the architecture of their cell nuclei", observes Prof. Wilczyński.