3-D tissue engineering: Novel technique brings researchers closer to viable organ implants
Researchers at the Institute of Bioengineering and Nanotechnology (IBN) have developed a simple method of organizing cells and their microenvironments in hydrogel fibers. Their unique technology provides a feasible template for assembling complex structures, such as liver and fat tissues, as described in their recent publication in Nature Communications.
According to IBN Executive Director Professor Jackie Y. Ying, "Our tissue engineering approach gives researchers great control and flexibility over the arrangement of individual cell types, making it possible to engineer prevascularized tissue constructs easily. This innovation brings us a step closer toward developing viable tissue or organ replacements."
IBN Team Leader and Principal Research Scientist, Dr Andrew Wan, elaborated, "Critical to the success of an implant is its ability to rapidly integrate with the patient's circulatory system. This is essential for the survival of cells within the implant, as it would ensure timely access to oxygen and essential nutrients, as well as the removal of metabolic waste products. Integration would also facilitate signaling between the cells and blood vessels, which is important for tissue development."
Tissues designed with pre-formed vascular networks are known to promote rapid vascular integration with the host. Generally, prevascularization has been achieved by seeding or encapsulating endothelial cells, which line the interior surfaces of blood vessels, with other cell types. In many of these approaches, the eventual distribution of vessels within a thick structure is reliant on in vitro cellular infiltration and self-organization of the cell mixture. These are slow processes, often leading to a non-uniform network of vessels within the tissue. As vascular self-assembly requires a large concentration of endothelial cells, this method also severely restricts the number of other cells that may be co-cultured.
Alternatively, scientists have attempted to direct the distribution of newly formed vessels via three-dimensional (3D) co-patterning of endothelial cells with other cell types in a hydrogel. This approach allows large concentrations of endothelial cells to be positioned in specific regions within the tissue, leaving the rest of the construct available for other cell types. The hydrogel also acts as a reservoir of nutrients for the encapsulated cells. However, co-patterning multiple cell types within a hydrogel is not easy. Conventional techniques, such as micromolding and organ printing, are limited by slow cell assembly, large volumes of cell suspension, complicated multi-step processes and expensive instruments. These factors also make it difficult to scale up the production of implantable 3D cell-patterned constructs. To date, these approaches have been unsuccessful in achieving vascularization and mass transport through thick engineered tissues.
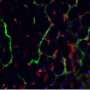
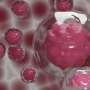
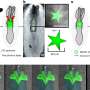
To overcome these limitations, IBN researchers have used interfacial polyelectrolyte complexation (IPC) fiber assembly, a unique cell patterning technology patented by IBN, to produce cell-laden hydrogel fibers under aqueous conditions at room temperature. Unlike other methods, IBN's novel technique allows researchers to incorporate different cell types separately into different fibers, and these cell-laden fibers may then be assembled into more complex constructs with hierarchical tissue structures. In addition, IBN researchers are able to tailor the microenvironment for each cell type for optimal functionality by incorporating the appropriate factors, e.g. proteins, into the fibers. Using IPC fiber assembly, the researchers have engineered an endothelial vessel network, as well as cell-patterned fat and liver tissue constructs, which have successfully integrated with the host circulatory system in a mouse model and produced vascularized tissues.
The IBN researchers are now working on applying and further developing their technology toward engineering functional tissues and clinical applications.
More information: Leong, M. et al. Patterned Prevascularized Tissue Constructs by Assembly of Polyelectrolyte Hydrogel Fibers, Nature Communications, (2013). DOI: 10.1038/ncomms3353.