Genetic test classifies kidney cancer
Singaporean researchers have developed a genetic test that can reliably identify different subtypes of a specific kind of kidney cancer. The test has the potential to improve treatment, as it can assess a patient's prognosis and likely response to therapy.
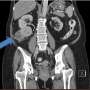
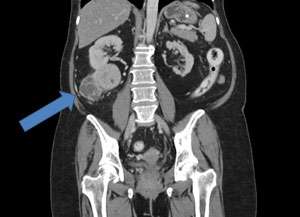
Clear cell renal cell carcinoma (ccRCC) is the most common type of kidney cancer (see image). The prognosis for patients with ccRCC who undergo surgery to remove a tumor is difficult to predict, however, since outcomes vary. The benefit of tyrosine kinase inhibitor (TKI) therapy—the most frequently used medication administered for ccRCC—also varies between patients. Evidence suggests that different outcomes could result from different subtypes of the disease; in which case, a test that identifies markers of these subtypes could improve treatment.
Min-Han Tan and co-workers from the A*STAR Institute of Bioengineering and Nanotechnology, Singapore General Hospital and National Cancer Centre Singapore aimed to develop a test that could identify genetic markers of different ccRCC subtypes. To look for relevant genes, they turned to genetic material preserved from 279 patients who had undergone surgery for ccRCC in the past 15 years.
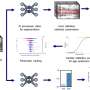
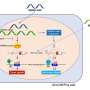
"We used microarrays to broadly survey all transcribed genes within the tissue sample," explains Tan. "We employed statistical methods to filter likely candidates, before using a separate method (quantitative real-time polymerase chain reaction) to determine which candidates had the best performance."
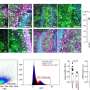
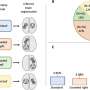
The team identified eight genes whose expression levels differed between two subtypes of ccRCC, allowing them to develop an assay based on these eight genes to test patients. They validated the test by using it on other groups of patients; the expression profiles of the eight genes proved to be a reliable predictor of a good or poor prognosis. Analysis of a subset of patients with ccRCC who received TKI therapy as well as surgery also showed that the assay can reliably predict whether patients will benefit from TKI therapy.
"This represents a real-world assay that could be used for both prognosis and prediction for ccRCC patients," explains Tan. "On a prognostic level, it allows an accessible way to determine good and poor outcomes in patients undergoing surgery. On a predictive level for TKIs, ours is the first study to associate biological subtypes with drug response."
Tan says that the assay could be used to improve surveillance strategies for patients with ccRCC, especially those with a poor prognosis, but that clinical trials are needed to determine the value of the assay in relation to various therapies.
More information: Choudhury, Y., Wei, X., Chu, Y.-H., Ng, L. G., Tan, H. S. et al. A multigene assay identifying distinct prognostic subtypes of clear cell renal cell carcinoma with differential response to tyrosine kinase inhibition. European Urology 67, 17–20 (2015). dx.doi.org/10.1016/j.eururo.2014.06.041